[English] 日本語
 Yorodumi
Yorodumi- PDB-1i11: SOLUTION STRUCTURE OF THE DNA BINDING DOMAIN, SOX-5 HMG BOX FROM MOUSE -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1i11 | ||||||
|---|---|---|---|---|---|---|---|
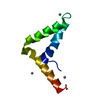
| Title | SOLUTION STRUCTURE OF THE DNA BINDING DOMAIN, SOX-5 HMG BOX FROM MOUSE | ||||||
 Components Components | TRANSCRIPTION FACTOR SOX-5 | ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / HMG BOX / DNA BENDING / DNA BINDING PROTEIN / HMG BOX / DNA BENDING /  DNA RECOGNITION / DNA RECOGNITION /  CHROMATIN / DNA SEQUENCE SPECIFIC / TESTIS DETERMINING. CHROMATIN / DNA SEQUENCE SPECIFIC / TESTIS DETERMINING. | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of mesenchymal stem cell differentiation / regulation of timing of neuron differentiation /  central nervous system neuron differentiation / positive regulation of cartilage development / asymmetric neuroblast division / positive regulation of chondrocyte differentiation / cartilage development / cartilage condensation / oligodendrocyte differentiation / central nervous system neuron differentiation / positive regulation of cartilage development / asymmetric neuroblast division / positive regulation of chondrocyte differentiation / cartilage development / cartilage condensation / oligodendrocyte differentiation /  cell fate commitment ...positive regulation of mesenchymal stem cell differentiation / regulation of timing of neuron differentiation / cell fate commitment ...positive regulation of mesenchymal stem cell differentiation / regulation of timing of neuron differentiation /  central nervous system neuron differentiation / positive regulation of cartilage development / asymmetric neuroblast division / positive regulation of chondrocyte differentiation / cartilage development / cartilage condensation / oligodendrocyte differentiation / central nervous system neuron differentiation / positive regulation of cartilage development / asymmetric neuroblast division / positive regulation of chondrocyte differentiation / cartilage development / cartilage condensation / oligodendrocyte differentiation /  cell fate commitment / cis-regulatory region sequence-specific DNA binding / chondrocyte differentiation / cellular response to transforming growth factor beta stimulus / in utero embryonic development / cell fate commitment / cis-regulatory region sequence-specific DNA binding / chondrocyte differentiation / cellular response to transforming growth factor beta stimulus / in utero embryonic development /  transcription regulator complex / transcription cis-regulatory region binding / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / negative regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II / transcription regulator complex / transcription cis-regulatory region binding / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / negative regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II /  nucleus nucleusSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  SOLUTION NMR / SIMULATING ANNEALING SOLUTION NMR / SIMULATING ANNEALING | ||||||
 Authors Authors | Cary, P.D. / Read, C.M. / Davis, B. / Driscoll, P.C. / Crane-Robinson, C. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2001 Journal: Protein Sci. / Year: 2001Title: Solution structure and backbone dynamics of the DNA-binding domain of mouse Sox-5. Authors: Cary, P.D. / Read, C.M. / Davis, B. / Driscoll, P.C. / Crane-Robinson, C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1i11.cif.gz 1i11.cif.gz | 700.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1i11.ent.gz pdb1i11.ent.gz | 586.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1i11.json.gz 1i11.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/i1/1i11 https://data.pdbj.org/pub/pdb/validation_reports/i1/1i11 ftp://data.pdbj.org/pub/pdb/validation_reports/i1/1i11 ftp://data.pdbj.org/pub/pdb/validation_reports/i1/1i11 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9825.412 Da / Num. of mol.: 1 / Fragment: HMG BOX Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Gene: SOX-5 / Organ: TESTIS Mus musculus (house mouse) / Gene: SOX-5 / Organ: TESTIS Testicle / Plasmid: PGEX-2T / Production host: Testicle / Plasmid: PGEX-2T / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21 (DE3) PLYSS / References: UniProt: P35710 Escherichia coli (E. coli) / Strain (production host): BL21 (DE3) PLYSS / References: UniProt: P35710 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||
| NMR details | Text: This Structure was determined using a combination of 2D and 3D NMR spectra. THE 30 STRUCTURES ARE ALIGNED OVER ALL BACKBONE ATOMS FOR RESIDUES 10-25 AND 32-43. MODEL 1 IS THE MINIMUM ENERGY AND ...Text: This Structure was determined using a combination of 2D and 3D NMR spectra. THE 30 STRUCTURES ARE ALIGNED OVER ALL BACKBONE ATOMS FOR RESIDUES 10-25 AND 32-43. MODEL 1 IS THE MINIMUM ENERGY AND REFERENCE STRUCTURE FOR THE OTHER 29 STRUCTURES. |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sample conditions | Ionic strength: 75 mM KPO4 / pH: 6.20 / Pressure: Ambient / Temperature: 298.00 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: SIMULATING ANNEALING / Software ordinal: 1 Details: Structures are based on a total of 1383 nonredundant distance NOE restraints, 61 dihedral angle restraints and 24 paired distance restraints for hydrogen bonds. | ||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations,structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 30 |
 Movie
Movie Controller
Controller









 PDBj
PDBj


