+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1f68 | ||||||
|---|---|---|---|---|---|---|---|
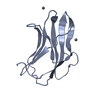
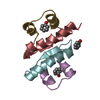
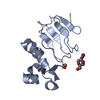
| Title | NMR SOLUTION STRUCTURE OF THE BROMODOMAIN FROM HUMAN GCN5 | ||||||
 Components Components | HISTONE ACETYLTRANSFERASE | ||||||
 Keywords Keywords |  TRANSFERASE / LEFT-HANDED FOUR-HELIX BUNDLE TRANSFERASE / LEFT-HANDED FOUR-HELIX BUNDLE | ||||||
| Function / homology |  Function and homology information Function and homology informationhistone succinyltransferase activity / peptidyl-lysine glutarylation / histone glutaryltransferase activity / metencephalon development / regulation of cartilage development / positive regulation of cell projection organization / : / histone H4K12 acetyltransferase activity / histone H3K9 acetyltransferase activity / positive regulation of cardiac muscle cell differentiation ...histone succinyltransferase activity / peptidyl-lysine glutarylation / histone glutaryltransferase activity / metencephalon development / regulation of cartilage development / positive regulation of cell projection organization / : / histone H4K12 acetyltransferase activity / histone H3K9 acetyltransferase activity / positive regulation of cardiac muscle cell differentiation / regulation of stem cell population maintenance /  regulation of bone development / regulation of regulatory T cell differentiation / negative regulation of centriole replication / transcription factor TFTC complex / telencephalon development / histone H3 acetyltransferase activity / internal peptidyl-lysine acetylation / histone H3K18 acetyltransferase activity / regulation of bone development / regulation of regulatory T cell differentiation / negative regulation of centriole replication / transcription factor TFTC complex / telencephalon development / histone H3 acetyltransferase activity / internal peptidyl-lysine acetylation / histone H3K18 acetyltransferase activity /  ATAC complex / ATAC complex /  Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / Cardiogenesis / SAGA complex / RUNX3 regulates NOTCH signaling / Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / Cardiogenesis / SAGA complex / RUNX3 regulates NOTCH signaling /  limb development / NOTCH4 Intracellular Domain Regulates Transcription / limb development / NOTCH4 Intracellular Domain Regulates Transcription /  regulation of T cell activation / NOTCH3 Intracellular Domain Regulates Transcription / regulation of tubulin deacetylation / peptide-lysine-N-acetyltransferase activity / midbrain development / intracellular distribution of mitochondria / Notch-HLH transcription pathway / Formation of WDR5-containing histone-modifying complexes / regulation of T cell activation / NOTCH3 Intracellular Domain Regulates Transcription / regulation of tubulin deacetylation / peptide-lysine-N-acetyltransferase activity / midbrain development / intracellular distribution of mitochondria / Notch-HLH transcription pathway / Formation of WDR5-containing histone-modifying complexes /  regulation of cell division / Formation of paraxial mesoderm / regulation of cell division / Formation of paraxial mesoderm /  regulation of RNA splicing / RNA Polymerase I Transcription Initiation / regulation of RNA splicing / RNA Polymerase I Transcription Initiation /  regulation of embryonic development / regulation of embryonic development /  histone acetyltransferase complex / negative regulation of gluconeogenesis / histone acetyltransferase complex / negative regulation of gluconeogenesis /  long-term memory / long-term memory /  regulation of DNA repair / regulation of DNA repair /  somitogenesis / somitogenesis /  histone acetyltransferase activity / positive regulation of gluconeogenesis / histone acetyltransferase activity / positive regulation of gluconeogenesis /  histone acetyltransferase / histone acetyltransferase /  Transferases; Acyltransferases; Transferring groups other than aminoacyl groups / response to nutrient levels / cellular response to nerve growth factor stimulus / neural tube closure / Transferases; Acyltransferases; Transferring groups other than aminoacyl groups / response to nutrient levels / cellular response to nerve growth factor stimulus / neural tube closure /  gluconeogenesis / positive regulation of cytokine production / gluconeogenesis / positive regulation of cytokine production /  regulation of synaptic plasticity / multicellular organism growth / regulation of synaptic plasticity / multicellular organism growth /  regulation of protein stability / B-WICH complex positively regulates rRNA expression / response to organic cyclic compound / NOTCH1 Intracellular Domain Regulates Transcription / regulation of protein stability / B-WICH complex positively regulates rRNA expression / response to organic cyclic compound / NOTCH1 Intracellular Domain Regulates Transcription /  mitotic spindle / Pre-NOTCH Transcription and Translation / Constitutive Signaling by NOTCH1 PEST Domain Mutants / Constitutive Signaling by NOTCH1 HD+PEST Domain Mutants / mitotic spindle / Pre-NOTCH Transcription and Translation / Constitutive Signaling by NOTCH1 PEST Domain Mutants / Constitutive Signaling by NOTCH1 HD+PEST Domain Mutants /  histone deacetylase binding / cellular response to tumor necrosis factor / histone deacetylase binding / cellular response to tumor necrosis factor /  heart development / HATs acetylate histones / fibroblast proliferation / heart development / HATs acetylate histones / fibroblast proliferation /  protein phosphatase binding / DNA-binding transcription factor binding / in utero embryonic development / protein phosphatase binding / DNA-binding transcription factor binding / in utero embryonic development /  transcription coactivator activity / transcription coactivator activity /  regulation of cell cycle / Ub-specific processing proteases / regulation of cell cycle / Ub-specific processing proteases /  chromatin remodeling / chromatin remodeling /  centrosome / centrosome /  chromatin binding / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / chromatin binding / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II /  extracellular space / extracellular space /  nucleoplasm / nucleoplasm /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / Torsion angle dynamics (DYANA) for generation of initial structures; Structure-mediated assignment of ambiguous experimental restraints (SANE); Iteration between DYANA, SANE; Simulated Annealing, Energy Minimization in AMBER using full restraint set. SOLUTION NMR / Torsion angle dynamics (DYANA) for generation of initial structures; Structure-mediated assignment of ambiguous experimental restraints (SANE); Iteration between DYANA, SANE; Simulated Annealing, Energy Minimization in AMBER using full restraint set. | ||||||
 Authors Authors | Wright, P.E. / Hudson, B.P. / Dyson, H.J. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2000 Journal: J.Mol.Biol. / Year: 2000Title: Solution structure and acetyl-lysine binding activity of the GCN5 bromodomain. Authors: Hudson, B.P. / Martinez-Yamout, M.A. / Dyson, H.J. / Wright, P.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1f68.cif.gz 1f68.cif.gz | 1001.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1f68.ent.gz pdb1f68.ent.gz | 850.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1f68.json.gz 1f68.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/f6/1f68 https://data.pdbj.org/pub/pdb/validation_reports/f6/1f68 ftp://data.pdbj.org/pub/pdb/validation_reports/f6/1f68 ftp://data.pdbj.org/pub/pdb/validation_reports/f6/1f68 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein |  / GCN5 / GCN5Mass: 12257.139 Da / Num. of mol.: 1 / Fragment: BROMODOMAIN / Mutation: P730G Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Organ: LIVER Homo sapiens (human) / Organ: LIVER / Plasmid: PET24A / Production host: / Plasmid: PET24A / Production host:   Escherichia coli (E. coli) / References: UniProt: Q92830, Escherichia coli (E. coli) / References: UniProt: Q92830,  histone acetyltransferase histone acetyltransferase |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||
| NMR details | Text: Backbone assignments from HNCACB & CBCA(CO)NH (sample 2); Sidechain assignments from C(CO)NH-TOCSY & H(CCO)NH-TOCSY (sample 2); Stereospecific assignments from 13C-{13CO} & 13C-{15N} spin-echo ...Text: Backbone assignments from HNCACB & CBCA(CO)NH (sample 2); Sidechain assignments from C(CO)NH-TOCSY & H(CCO)NH-TOCSY (sample 2); Stereospecific assignments from 13C-{13CO} & 13C-{15N} spin-echo difference CT-HSQC (sample 3); Aromatic assignments from (HB)CB(CGCD)HD, (HB)CB(CGCDCE)HE, & (HC)C(C)CH-TOCSY (sample 3) |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions |
| ||||||||||||||||||||
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: Torsion angle dynamics (DYANA) for generation of initial structures; Structure-mediated assignment of ambiguous experimental restraints (SANE); Iteration between DYANA, SANE; Simulated ...Method: Torsion angle dynamics (DYANA) for generation of initial structures; Structure-mediated assignment of ambiguous experimental restraints (SANE); Iteration between DYANA, SANE; Simulated Annealing, Energy Minimization in AMBER using full restraint set. Software ordinal: 1 Details: Structures are based upon 2232 NOE-derived distance restraints (365 long-range, 379 medium-range, 403 sequential, 1085 intraresidue, and 426 ambiguous), 155 torsion angle restraints, and 47 ...Details: Structures are based upon 2232 NOE-derived distance restraints (365 long-range, 379 medium-range, 403 sequential, 1085 intraresidue, and 426 ambiguous), 155 torsion angle restraints, and 47 stereospecific assignments. | ||||||||||||||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: NO DISTANCE OR ANGLE VIOLATIONS GREATER THAN 0.15 A OR 5 DEGREES Conformers calculated total number: 185 / Conformers submitted total number: 30 |
 Movie
Movie Controller
Controller










 PDBj
PDBj



