+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1e0n | ||||||
|---|---|---|---|---|---|---|---|
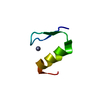
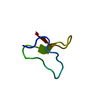
| Title | YJQ8WW domain from Saccharomyces cerevisae | ||||||
 Components Components | HYPOTHETICAL PROTEIN | ||||||
 Keywords Keywords | YJQ8WW DOMAIN /  WW DOMAIN / SACCHAROMYCES CEREVISAE / YJQ8 PROTEIN WW DOMAIN / SACCHAROMYCES CEREVISAE / YJQ8 PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of antisense RNA transcription / ascospore formation / negative regulation of reciprocal meiotic recombination / [histone H3]-lysine36 N-trimethyltransferase / histone H3K36 trimethyltransferase activity / DNA-templated transcription elongation / histone H3K36 methyltransferase activity / regulation of DNA-templated DNA replication initiation / DNA-templated transcription termination /  methylation ...negative regulation of antisense RNA transcription / ascospore formation / negative regulation of reciprocal meiotic recombination / [histone H3]-lysine36 N-trimethyltransferase / histone H3K36 trimethyltransferase activity / DNA-templated transcription elongation / histone H3K36 methyltransferase activity / regulation of DNA-templated DNA replication initiation / DNA-templated transcription termination / methylation ...negative regulation of antisense RNA transcription / ascospore formation / negative regulation of reciprocal meiotic recombination / [histone H3]-lysine36 N-trimethyltransferase / histone H3K36 trimethyltransferase activity / DNA-templated transcription elongation / histone H3K36 methyltransferase activity / regulation of DNA-templated DNA replication initiation / DNA-templated transcription termination /  methylation / methylation /  chromatin / regulation of DNA-templated transcription / chromatin / regulation of DNA-templated transcription /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   SACCHAROMYCES CEREVISIAE (brewer's yeast) SACCHAROMYCES CEREVISIAE (brewer's yeast) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Macias, M.J. / Gervais, V. / Civera, C. / Oschkinat, H. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2000 Journal: Nat.Struct.Biol. / Year: 2000Title: Structural Analysis of Ww Domains and Design of a Ww Prototype Authors: Macias, M.J. / Gervais, V. / Civera, C. / Oschkinat, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1e0n.cif.gz 1e0n.cif.gz | 46.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1e0n.ent.gz pdb1e0n.ent.gz | 30 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1e0n.json.gz 1e0n.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/e0/1e0n https://data.pdbj.org/pub/pdb/validation_reports/e0/1e0n ftp://data.pdbj.org/pub/pdb/validation_reports/e0/1e0n ftp://data.pdbj.org/pub/pdb/validation_reports/e0/1e0n | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1e0lC  1e0mC C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide |  Mass: 3256.645 Da / Num. of mol.: 1 / Fragment: DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)   SACCHAROMYCES CEREVISIAE (brewer's yeast) SACCHAROMYCES CEREVISIAE (brewer's yeast)Production host:   ESCHERICHIA COLI (E. coli) / References: UniProt: P46995*PLUS ESCHERICHIA COLI (E. coli) / References: UniProt: P46995*PLUS |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||
| NMR details | Text: THE STRUCTURE WAS DETERMINED USING HOMONUCLEAR NMR |
- Sample preparation
Sample preparation
| Sample conditions | pH: 6.5 / Temperature: 285 K |
|---|
-NMR measurement
| NMR spectrometer | Type: Bruker DRX / Manufacturer: Bruker / Model : DRX / Field strength: 600 MHz : DRX / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 | |||||||||
| NMR ensemble | Conformer selection criteria: LOWEST ENERGY / Conformers calculated total number: 20 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller









 PDBj
PDBj
