[English] 日本語
 Yorodumi
Yorodumi- PDB-1dy9: Inhibition of the Hepatitis C Virus NS3/4A Protease. The Crystal ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1dy9 | ||||||
|---|---|---|---|---|---|---|---|
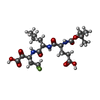
| Title | Inhibition of the Hepatitis C Virus NS3/4A Protease. The Crystal Structures of Two Protease-Inhibitor Complexes (inhibitor I) | ||||||
 Components Components |
| ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE INHIBITOR /  SERINE PROTEASE / NS3 / SERINE PROTEASE / NS3 /  NS4A / NS4A /  HEPATITIS C VIRUS / PROTEASE INHIBITION / HYDROLASE-HYDROLASE INHIBITOR COMPLEX HEPATITIS C VIRUS / PROTEASE INHIBITION / HYDROLASE-HYDROLASE INHIBITOR COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationRNA stabilization / RNA folding chaperone / DNA/DNA annealing activity / RNA strand annealing activity /  hepacivirin / host cell mitochondrial membrane / host cell lipid droplet / symbiont-mediated suppression of host TRAF-mediated signal transduction / transformation of host cell by virus / symbiont-mediated perturbation of host cell cycle G1/S transition checkpoint ...RNA stabilization / RNA folding chaperone / DNA/DNA annealing activity / RNA strand annealing activity / hepacivirin / host cell mitochondrial membrane / host cell lipid droplet / symbiont-mediated suppression of host TRAF-mediated signal transduction / transformation of host cell by virus / symbiont-mediated perturbation of host cell cycle G1/S transition checkpoint ...RNA stabilization / RNA folding chaperone / DNA/DNA annealing activity / RNA strand annealing activity /  hepacivirin / host cell mitochondrial membrane / host cell lipid droplet / symbiont-mediated suppression of host TRAF-mediated signal transduction / transformation of host cell by virus / symbiont-mediated perturbation of host cell cycle G1/S transition checkpoint / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / hepacivirin / host cell mitochondrial membrane / host cell lipid droplet / symbiont-mediated suppression of host TRAF-mediated signal transduction / transformation of host cell by virus / symbiont-mediated perturbation of host cell cycle G1/S transition checkpoint / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity /  protein-DNA complex / protein-DNA complex /  : / nucleoside-triphosphate phosphatase / protein complex oligomerization / monoatomic ion channel activity / viral nucleocapsid / clathrin-dependent endocytosis of virus by host cell / : / nucleoside-triphosphate phosphatase / protein complex oligomerization / monoatomic ion channel activity / viral nucleocapsid / clathrin-dependent endocytosis of virus by host cell /  Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases / Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases /  RNA helicase activity / host cell endoplasmic reticulum membrane / host cell perinuclear region of cytoplasm / RNA helicase activity / host cell endoplasmic reticulum membrane / host cell perinuclear region of cytoplasm /  RNA helicase / induction by virus of host autophagy / RNA helicase / induction by virus of host autophagy /  ribonucleoprotein complex / ribonucleoprotein complex /  RNA-directed RNA polymerase / viral RNA genome replication / cysteine-type endopeptidase activity / RNA-directed RNA polymerase / viral RNA genome replication / cysteine-type endopeptidase activity /  RNA-dependent RNA polymerase activity / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / RNA-dependent RNA polymerase activity / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane /  viral envelope / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / host cell nucleus / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / viral envelope / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / host cell nucleus / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity /  ATP hydrolysis activity / ATP hydrolysis activity /  proteolysis / proteolysis /  DNA binding / DNA binding /  RNA binding / zinc ion binding / RNA binding / zinc ion binding /  ATP binding / ATP binding /  membrane membraneSimilarity search - Function | ||||||
| Biological species |  HEPATITIS C VIRUS HEPATITIS C VIRUS | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å MOLECULAR REPLACEMENT / Resolution: 2.1 Å | ||||||
 Authors Authors | Di Marco, S. / Rizzi, M. / Volpari, C. / Walsh, M. / Narjes, F. / Colarusso, S. / De Francesco, R. / Matassa, V.G. / Sollazzo, M. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2000 Journal: J.Biol.Chem. / Year: 2000Title: Inhibition of the Hepatitis C Virus Ns3/4A Protease the Crystal Structures of Two Protease-Inhibitor Complexes Authors: Di Marco, S. / Rizzi, M. / Volpari, C. / Walsh, M. / Narjes, F. / Colarusso, S. / De Francesco, R. / Matassa, V.G. / Sollazzo, M. | ||||||
| History |
| ||||||
| Remark 700 | SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS AC AND BC IN EACH CHAIN ON SHEET RECORDS ... SHEET DETERMINATION METHOD: DSSP THE SHEETS PRESENTED AS AC AND BC IN EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 6-STRANDED BARREL THIS IS REPRESENTED BY A 7-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1dy9.cif.gz 1dy9.cif.gz | 91 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1dy9.ent.gz pdb1dy9.ent.gz | 67.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1dy9.json.gz 1dy9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dy/1dy9 https://data.pdbj.org/pub/pdb/validation_reports/dy/1dy9 ftp://data.pdbj.org/pub/pdb/validation_reports/dy/1dy9 ftp://data.pdbj.org/pub/pdb/validation_reports/dy/1dy9 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1dxpC  1dy8C  1dy7S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 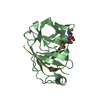
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (0.3724, 0.92807, -0.0026), Vector  : : |
- Components
Components
| #1: Protein | Mass: 19749.553 Da / Num. of mol.: 2 / Fragment: PROTEASE Source method: isolated from a genetically manipulated source Source: (gene. exp.)   HEPATITIS C VIRUS (ISOLATE TAIWAN) / Strain: ISOLATE TAIWAN / Description: CDNA OF HEPATITIS C VIRUS / Gene: HCV / Plasmid: PT7-7 / Production host: HEPATITIS C VIRUS (ISOLATE TAIWAN) / Strain: ISOLATE TAIWAN / Description: CDNA OF HEPATITIS C VIRUS / Gene: HCV / Plasmid: PT7-7 / Production host:   ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3)References: UniProt: Q81755, UniProt: P26662*PLUS,  Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases#2: Protein/peptide | Mass: 1686.097 Da / Num. of mol.: 2 / Fragment: RESIDUES 956-967 / Mutation: YES / Source method: obtained synthetically / Source: (synth.)  HEPATITIS C VIRUS / References: UniProt: Q81755, UniProt: P26662*PLUS HEPATITIS C VIRUS / References: UniProt: Q81755, UniProt: P26662*PLUS#3: Chemical | #4: Chemical | ChemComp-ZN / | #5: Water | ChemComp-HOH / |  Water WaterSequence details | GENOME POLYPROTEIN, ID POLG_HCVJA_7, PROTEASE NS3 GENOME POLYPROTEIN, ID POLG_HCVJA_8, ...GENOME POLYPROTEI | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.37 Å3/Da / Density % sol: 47.68 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Method: vapor diffusion / pH: 5.1 Details: THE NS3 PROTEIN (1MG/ML) WAS INCUBATE AT 4C WITH THE NS4A COFACTOR PEPTIDE, CONTAINING A SOLUBIZING LYSINE TAG AT ITS N- AND C-TERMINI(KGSVVIVGRIILSGRK), AT A MOLAR RATIO OF 1:2 AND ...Details: THE NS3 PROTEIN (1MG/ML) WAS INCUBATE AT 4C WITH THE NS4A COFACTOR PEPTIDE, CONTAINING A SOLUBIZING LYSINE TAG AT ITS N- AND C-TERMINI(KGSVVIVGRIILSGRK), AT A MOLAR RATIO OF 1:2 AND CONCENTRATED TO 290 MICROMOLAR. NS3J/4A CRYSTALS, WITH A MAXIMUM SIZE OF 0.6 X 0.3 X 0.2 MM**3, WERE OBTAINED BY BOTH HANGING- AND SITTING-DROP VAPOUR DIFFUSION METHODS AFTER TWO WEEKS AT ROOM TEMPERATURE, WITH 3.4 M NACL, 10MM DTT, 0.1 M CITRATE BUFFER PH 5.1. THE TERNARY COMPLEX WITH INHIBITOR WAS PREPARED BY ADDING 5.0 MM OF INIBITOR TO CRYSTALS THAT WERE STABILISED IN 4.5 M NACL, 10 MM DTT, 0.1 M CITRATE BUFFER, PH 5.1. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-3 / Wavelength: 0.936 / Beamline: ID14-3 / Wavelength: 0.936 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Feb 15, 1999 / Details: TOROIDAL MIRROR |
| Radiation | Monochromator: SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.936 Å / Relative weight: 1 : 0.936 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→20 Å / Num. obs: 23960 / % possible obs: 98.2 % / Observed criterion σ(I): -3 / Redundancy: 2.8 % / Biso Wilson estimate: 37.4 Å2 / Rsym value: 0.035 / Net I/σ(I): 32 |
| Reflection shell | Resolution: 2.1→2.14 Å / Redundancy: 2.1 % / Mean I/σ(I) obs: 3.2 / Rsym value: 0.15 / % possible all: 98.4 |
| Reflection | *PLUS Num. measured all: 66924 / Rmerge(I) obs: 0.035 |
| Reflection shell | *PLUS % possible obs: 98.4 % / Rmerge(I) obs: 0.15 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1DY7 Resolution: 2.1→20 Å / SU B: 3.62 / SU ML: 0.097 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.22 / ESU R Free: 0.21
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 43.5 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj