[English] 日本語
 Yorodumi
Yorodumi- PDB-1atd: HIGH-RESOLUTION STRUCTURE OF ASCARIS TRYPSIN INHIBITOR IN SOLUTIO... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1atd | ||||||
|---|---|---|---|---|---|---|---|
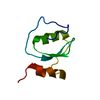
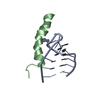
| Title | HIGH-RESOLUTION STRUCTURE OF ASCARIS TRYPSIN INHIBITOR IN SOLUTION: DIRECT EVIDENCE FOR A PH INDUCED CONFORMATIONAL TRANSITION IN THE REACTIVE SITE | ||||||
 Components Components | ASCARIS TRYPSIN INHIBITOR | ||||||
 Keywords Keywords | PROTEINASE INHIBITOR(TRYPSIN) | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |   Ascaris suum (pig roundworm) Ascaris suum (pig roundworm) | ||||||
| Method |  SOLUTION NMR SOLUTION NMR | ||||||
 Authors Authors | Clore, G.M. / Grasberger, B.L. / Gronenborn, A.M. | ||||||
 Citation Citation |  Journal: Structure / Year: 1994 Journal: Structure / Year: 1994Title: High-resolution structure of Ascaris trypsin inhibitor in solution: direct evidence for a pH-induced conformational transition in the reactive site. Authors: Grasberger, B.L. / Clore, G.M. / Gronenborn, A.M. #1:  Journal: Biochemistry / Year: 1990 Journal: Biochemistry / Year: 1990Title: Sequential Resonance Assignment and Secondary Structure Determination of the Ascaris Trypsin Inhibitor, a Member of a Novel Class of Proteinase Inhibitors Authors: Gronenborn, A.M. / Nilges, M. / Peanasky, R.J. / Clore, G.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1atd.cif.gz 1atd.cif.gz | 564.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1atd.ent.gz pdb1atd.ent.gz | 490.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1atd.json.gz 1atd.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/at/1atd https://data.pdbj.org/pub/pdb/validation_reports/at/1atd ftp://data.pdbj.org/pub/pdb/validation_reports/at/1atd ftp://data.pdbj.org/pub/pdb/validation_reports/at/1atd | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 6807.853 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Ascaris suum (pig roundworm) / References: UniProt: P19398 Ascaris suum (pig roundworm) / References: UniProt: P19398 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
Crystal grow | *PLUS Method: other / Details: NMR |
|---|
- Processing
Processing
| Refinement | Software ordinal: 1 Details: THE 3D STRUCTURE OF THE PH 4.75 FORM OF THE ASCARIS TRYPSIN INHIBITOR IN SOLUTION BY NMR IS BASED ON 1078 EXPERIMENTAL RESTRAINTS COMPRISING: 43 SHORT RANGE (1 < |I-J| 5) INTERRESIDUE ...Details: THE 3D STRUCTURE OF THE PH 4.75 FORM OF THE ASCARIS TRYPSIN INHIBITOR IN SOLUTION BY NMR IS BASED ON 1078 EXPERIMENTAL RESTRAINTS COMPRISING: 43 SHORT RANGE (1 < |I-J| <=5) AND 218 LONG RANGE (|I-J|>5) INTERRESIDUE INTERPROTON DISTANCE RESTRAINTS, 320 INTRARESIDUE INTERPROTON DISTANCE RESTRAINTS, 48 DISTANCE RESTRAINTS FOR 4 HYDROGEN BONDS, AND 59 PHI, 49 PSI AND 41 CHI1 TORSION ANGLE RESTRAINTS. A COMPLETE LIST OF EXPERIMENTAL RESTRAINTS HAS BEEN DEPOSITED WITH THE BROOKHAVEN DATA BANK. THE STRUCTURES ARE CALCULATED USING THE HYBRID METRIC MATRIX DISTANCE GEOMETRY-DYNAMICAL SIMULATED ANNEALING METHOD DESCRIBED BY: NILGES, M., CLORE, G.M. & GRONENBORN, A.M. (1988) FEBS LETT 29, 317-324 ALL STRUCTURAL STATISTICS ARE GIVEN IN REFERENCE 1. THE RESTRAINED MINIMIZED AVERAGE STRUCTURE (SA)R IS PRESENTED IN PROTEIN DATA BANK ENTRY 1ATA. THIS IS OBTAINED BY FIRST AVERAGING THE COORDINATES OF THE INDIVIDUAL 32 DYNAMICAL SIMULATED ANNEALING SA STRUCTURES BEST FITTED TO RESIDUES 5 - 60, AND SUBJECTING THE RESULTING COORDINATES TO RESTRAINED MINIMIZATION. THE QUANTITY PRESENTED IN COLUMNS 61 - 66 IN ENTRY 1ATA (THE B VALUE FIELD IN X-RAY STRUCTURES) GIVES THE AVERAGE RMS DIFFERENCE BETWEEN THE INDIVIDUAL SA STRUCTURES AND THE MEAN STRUCTURE. THE QUANTITIES IN THIS FIELD OF THE INDIVIDUAL STRUCTURES IN THIS ENTRY HAVE NO MEANING. |
|---|---|
| NMR ensemble | Conformers submitted total number: 32 |
 Movie
Movie Controller
Controller













 PDBj
PDBj