[English] 日本語
 Yorodumi
Yorodumi- EMDB-4625: E. coli ClpB (DWB and K476C mutant) bound to casein - state KC-2 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4625 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli ClpB (DWB and K476C mutant) bound to casein - state KC-2 | ||||||||||||
 Map data Map data | ClpB-DWB-K476C bound to casein in presence of ATPgammaS - state KC-2 | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |   Escherichia coli (E. coli) / Escherichia coli (E. coli) /   Bos taurus (cattle) Bos taurus (cattle) | ||||||||||||
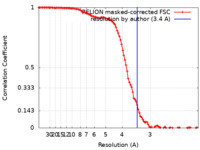
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.4 Å cryo EM / Resolution: 3.4 Å | ||||||||||||
 Authors Authors | Deville C / Saibil HR | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2019 Journal: Cell Rep / Year: 2019Title: Two-Step Activation Mechanism of the ClpB Disaggregase for Sequential Substrate Threading by the Main ATPase Motor. Authors: Célia Deville / Kamila Franke / Axel Mogk / Bernd Bukau / Helen R Saibil /   Abstract: AAA+ proteins form asymmetric hexameric rings that hydrolyze ATP and thread substrate proteins through a central channel via mobile substrate-binding pore loops. Understanding how ATPase and ...AAA+ proteins form asymmetric hexameric rings that hydrolyze ATP and thread substrate proteins through a central channel via mobile substrate-binding pore loops. Understanding how ATPase and threading activities are regulated and intertwined is key to understanding the AAA+ protein mechanism. We studied the disaggregase ClpB, which contains tandem ATPase domains (AAA1, AAA2) and shifts between low and high ATPase and threading activities. Coiled-coil M-domains repress ClpB activity by encircling the AAA1 ring. Here, we determine the mechanism of ClpB activation by comparing ATPase mechanisms and cryo-EM structures of ClpB wild-type and a constitutively active ClpB M-domain mutant. We show that ClpB activation reduces ATPase cooperativity and induces a sequential mode of ATP hydrolysis in the AAA2 ring, the main ATPase motor. AAA1 and AAA2 rings do not work synchronously but in alternating cycles. This ensures high grip, enabling substrate threading via a processive, rope-climbing mechanism. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4625.map.gz emd_4625.map.gz | 85.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4625-v30.xml emd-4625-v30.xml emd-4625.xml emd-4625.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4625_fsc.xml emd_4625_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_4625.png emd_4625.png | 88.2 KB | ||
| Masks |  emd_4625_msk_1.map emd_4625_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Others |  emd_4625_half_map_1.map.gz emd_4625_half_map_1.map.gz emd_4625_half_map_2.map.gz emd_4625_half_map_2.map.gz | 29.7 MB 29.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4625 http://ftp.pdbj.org/pub/emdb/structures/EMD-4625 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4625 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4625 | HTTPS FTP |
-Related structure data
| Related structure data |  4621C  4622C  4623C  4624C  4626C  4627C  4940C  4941C  4942C  6qs4C  6qs6C  6qs7C  6qs8C  6rn2C  6rn3C  6rn4C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4625.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4625.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ClpB-DWB-K476C bound to casein in presence of ATPgammaS - state KC-2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_4625_msk_1.map emd_4625_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_4625_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_4625_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ClpB-DWB-K476C bound to casein in presence of ATPgammaS
| Entire | Name: ClpB-DWB-K476C bound to casein in presence of ATPgammaS |
|---|---|
| Components |
|
-Supramolecule #1: ClpB-DWB-K476C bound to casein in presence of ATPgammaS
| Supramolecule | Name: ClpB-DWB-K476C bound to casein in presence of ATPgammaS type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Molecular weight | Theoretical: 570 KDa |
-Macromolecule #1: ClpB
| Macromolecule | Name: ClpB / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MRLDRLTNKF QLALADAQSL ALGHDNQFIE PLHLMSALLN QEGGSVSPLL TSAGINAGQL RTDINQALN RLPQVEGTGG DVQPSQDLVR VLNLCDKLAQ KRGDNFISSE LFVLAALESR G TLADILKA AGATTANITQ AIEQMRGGES VNDQGAEDQR QALKKYTIDL ...String: MRLDRLTNKF QLALADAQSL ALGHDNQFIE PLHLMSALLN QEGGSVSPLL TSAGINAGQL RTDINQALN RLPQVEGTGG DVQPSQDLVR VLNLCDKLAQ KRGDNFISSE LFVLAALESR G TLADILKA AGATTANITQ AIEQMRGGES VNDQGAEDQR QALKKYTIDL TERAEQGKLD PV IGRDEEI RRTIQVLQRR TKNNPVLIGE PGVGKTAIVE GLAQRIINGE VPEGLKGRRV LAL DMGALV AGAKYRGEFE ERLKGVLNDL AKQEGNVILF IDALHTMVGA GKADGAMDAG NMLK PALAR GELHCVGATT LDEYRQYIEK DAALERRFQK VFVAEPSVED TIAILRGLKE RYELH HHVQ ITDPAIVAAA TLSHRYIADR QLPDKAIDLI DEAASSIRMQ IDSKPEELDR LDRRII QLK LEQQALMKES DEASKKRLDM LNEELSDKER QYSELEEEWK AEKASLSGTQ TICAELE QA KIAIEQARRV GDLARMSELQ YGKIPELEKQ LEAATQLEGK TMRLLRNKVT DAEIAEVL A RWTGIPVSRM MESEREKLLR MEQELHHRVI GQNEAVDAVS NAIRRSRAGL ADPNRPIGS FLFLGPTGVG KTELCKALAN FMFDSDEAMV RIDMSEFMEK HSVSRLVGAP PGYVGYEEGG YLTEAVRRR PYSVILLDAV EKAHPDVFNI LLQVLDDGRL TDGQGRTVDF RNTVVIMTSN L GSDLIQER FGELDYAHMK ELVLGVVSHN FRPEFINRID EVVVFHPLGE QHIASIAQIQ LK RLYKRLE ERGYEIHISD EALKLLSENG YDPVYGARPL KRAIQQQIEN PLAQQILSGE LVP GKVIRL EVNEDRIVAV Q |
-Macromolecule #2: casein
| Macromolecule | Name: casein / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Bos taurus (cattle) Bos taurus (cattle) |
| Sequence | String: MKLLILTCLV AVALARPKHP IKHQGLPQEV LNENLLRFFV APFPEVFGKE KVNELSKDIG SESTEDQAM EDIKQMEAES ISSSEEIVPN SVEQKHIQKE DVPSERYLGY LEQLLRLKKY K VPQLEIVP NSAEERLHSM KEGIHAQQKE PMIGVNQELA YFYPELFRQF ...String: MKLLILTCLV AVALARPKHP IKHQGLPQEV LNENLLRFFV APFPEVFGKE KVNELSKDIG SESTEDQAM EDIKQMEAES ISSSEEIVPN SVEQKHIQKE DVPSERYLGY LEQLLRLKKY K VPQLEIVP NSAEERLHSM KEGIHAQQKE PMIGVNQELA YFYPELFRQF YQLDAYPSGA WY YVPLGTQ YTDAPSFSDI PNPIGSENSE KTTMPLW |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.6 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
| Grid | Model: Quantifoil, UltrAuFoil / Material: GOLD / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm |
| Specialist optics | Energy filter - Name: GIF Bioquantum |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 6060 / Average electron dose: 1.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X