+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human alpha-2/delta-1 without mirogabalin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of membrane repolarization during action potential / positive regulation of high voltage-gated calcium channel activity / calcium ion transmembrane transport via high voltage-gated calcium channel / membrane depolarization during bundle of His cell action potential /  L-type voltage-gated calcium channel complex / cardiac muscle cell action potential involved in contraction / regulation of ventricular cardiac muscle cell membrane repolarization / calcium ion transport into cytosol / regulation of calcium ion transmembrane transport via high voltage-gated calcium channel / L-type voltage-gated calcium channel complex / cardiac muscle cell action potential involved in contraction / regulation of ventricular cardiac muscle cell membrane repolarization / calcium ion transport into cytosol / regulation of calcium ion transmembrane transport via high voltage-gated calcium channel /  voltage-gated calcium channel complex ...regulation of membrane repolarization during action potential / positive regulation of high voltage-gated calcium channel activity / calcium ion transmembrane transport via high voltage-gated calcium channel / membrane depolarization during bundle of His cell action potential / voltage-gated calcium channel complex ...regulation of membrane repolarization during action potential / positive regulation of high voltage-gated calcium channel activity / calcium ion transmembrane transport via high voltage-gated calcium channel / membrane depolarization during bundle of His cell action potential /  L-type voltage-gated calcium channel complex / cardiac muscle cell action potential involved in contraction / regulation of ventricular cardiac muscle cell membrane repolarization / calcium ion transport into cytosol / regulation of calcium ion transmembrane transport via high voltage-gated calcium channel / L-type voltage-gated calcium channel complex / cardiac muscle cell action potential involved in contraction / regulation of ventricular cardiac muscle cell membrane repolarization / calcium ion transport into cytosol / regulation of calcium ion transmembrane transport via high voltage-gated calcium channel /  voltage-gated calcium channel complex / neuronal dense core vesicle / regulation of heart rate by cardiac conduction / regulation of calcium ion transport / calcium ion import across plasma membrane / voltage-gated calcium channel complex / neuronal dense core vesicle / regulation of heart rate by cardiac conduction / regulation of calcium ion transport / calcium ion import across plasma membrane /  voltage-gated calcium channel activity / voltage-gated calcium channel activity /  sarcoplasmic reticulum / cellular response to amyloid-beta / calcium ion transport / extracellular exosome / sarcoplasmic reticulum / cellular response to amyloid-beta / calcium ion transport / extracellular exosome /  metal ion binding / metal ion binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
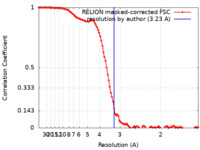
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.23 Å cryo EM / Resolution: 3.23 Å | |||||||||
 Authors Authors | Kozai D / Numoto N / Fujiyoshi Y | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2023 Journal: J Mol Biol / Year: 2023Title: Recognition Mechanism of a Novel Gabapentinoid Drug, Mirogabalin, for Recombinant Human αδ1, a Voltage-Gated Calcium Channel Subunit. Authors: Daisuke Kozai / Nobutaka Numoto / Kouki Nishikawa / Akiko Kamegawa / Shohei Kawasaki / Yoko Hiroaki / Katsumasa Irie / Atsunori Oshima / Hiroyuki Hanzawa / Kousei Shimada / Yutaka Kitano / Yoshinori Fujiyoshi /  Abstract: Mirogabalin is a novel gabapentinoid drug with a hydrophobic bicyclo substituent on the γ-aminobutyric acid moiety that targets the voltage-gated calcium channel subunit αδ1. Here, to reveal the ...Mirogabalin is a novel gabapentinoid drug with a hydrophobic bicyclo substituent on the γ-aminobutyric acid moiety that targets the voltage-gated calcium channel subunit αδ1. Here, to reveal the mirogabalin recognition mechanisms of αδ1, we present structures of recombinant human αδ1 with and without mirogabalin analyzed by cryo-electron microscopy. These structures show the binding of mirogabalin to the previously reported gabapentinoid binding site, which is the extracellular dCache_1 domain containing a conserved amino acid binding motif. A slight conformational change occurs around the residues positioned close to the hydrophobic group of mirogabalin. Mutagenesis binding assays identified that residues in the hydrophobic interaction region, in addition to several amino acid binding motif residues around the amino and carboxyl groups of mirogabalin, are critical for mirogabalin binding. The A215L mutation introduced to decrease the hydrophobic pocket volume predictably suppressed mirogabalin binding and promoted the binding of another ligand, L-Leu, with a smaller hydrophobic substituent than mirogabalin. Alterations of residues in the hydrophobic interaction region of αδ1 to those of the αδ2, αδ3, and αδ4 isoforms, of which αδ3 and αδ4 are gabapentin-insensitive, suppressed the binding of mirogabalin. These results support the importance of hydrophobic interactions in αδ1 ligand recognition. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35400.map.gz emd_35400.map.gz | 23.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35400-v30.xml emd-35400-v30.xml emd-35400.xml emd-35400.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35400_fsc.xml emd_35400_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_35400.png emd_35400.png | 136.5 KB | ||
| Masks |  emd_35400_msk_1.map emd_35400_msk_1.map | 25.3 MB |  Mask map Mask map | |
| Others |  emd_35400_half_map_1.map.gz emd_35400_half_map_1.map.gz emd_35400_half_map_2.map.gz emd_35400_half_map_2.map.gz | 23.5 MB 23.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35400 http://ftp.pdbj.org/pub/emdb/structures/EMD-35400 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35400 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35400 | HTTPS FTP |
-Related structure data
| Related structure data |  8if4MC  8if3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35400.map.gz / Format: CCP4 / Size: 25.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35400.map.gz / Format: CCP4 / Size: 25.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.765 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_35400_msk_1.map emd_35400_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
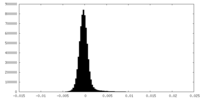
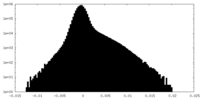
| Density Histograms |
-Half map: #1
| File | emd_35400_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_35400_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Voltage-dependent calcium channel subunit alpha-2/delta-1
| Entire | Name: Voltage-dependent calcium channel subunit alpha-2/delta-1 Voltage-gated calcium channel Voltage-gated calcium channel |
|---|---|
| Components |
|
-Supramolecule #1: Voltage-dependent calcium channel subunit alpha-2/delta-1
| Supramolecule | Name: Voltage-dependent calcium channel subunit alpha-2/delta-1 type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 140 KDa |
-Macromolecule #1: Voltage-dependent calcium channel subunit alpha-2/delta-1
| Macromolecule | Name: Voltage-dependent calcium channel subunit alpha-2/delta-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 124.413062 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAAGCLLALT LTLFQSLLIG PSSEEPFPSA VTIKSWVDKM QEDLVTLAKT ASGVNQLVDI YEKYQDLYTV EPNNARQLVE IAARDIEKL LSNRSKALVR LALEAEKVQA AHQWREDFAS NEVVYYNAKD DLDPEKNDSE PGSQRIKPVF IEDANFGRQI S YQHAAVHI ...String: MAAGCLLALT LTLFQSLLIG PSSEEPFPSA VTIKSWVDKM QEDLVTLAKT ASGVNQLVDI YEKYQDLYTV EPNNARQLVE IAARDIEKL LSNRSKALVR LALEAEKVQA AHQWREDFAS NEVVYYNAKD DLDPEKNDSE PGSQRIKPVF IEDANFGRQI S YQHAAVHI PTDIYEGSTI VLNELNWTSA LDEVFKKNRE EDPSLLWQVF GSATGLARYY PASPWVDNSR TPNKIDLYDV RR RPWYIQG AASPKDMLIL VDVSGSVSGL TLKLIRTSVS EMLETLSDDD FVNVASFNSN AQDVSCFQHL VQANVRNKKV LKD AVNNIT AKGITDYKKG FSFAFEQLLN YNVSRANCNK IIMLFTDGGE ERAQEIFNKY NKDKKVRVFT FSVGQHNYDR GPIQ WMACE NKGYYYEIPS IGAIRINTQE YLDVLGRPMV LAGDKAKQVQ WTNVYLDALE LGLVITGTLP VFNITGQFEN KTNLK NQLI LGVMGVDVSL EDIKRLTPRF TLCPNGYYFA IDPNGYVLLH PNLQPKNPKS QEPVTLDFLD AELENDIKVE IRNKMI DGE SGEKTFRTLV KSQDERYIDK GNRTYTWTPV NGTDYSLALV LPTYSFYYIK AKLEETITQA RSKKGKMKDS ETLKPDN FE ESGYTFIAPR DYCNDLKISD NNTEFLLNFN EFIDRHHHHH HHHKTPNNPS CNADLINRVL LDAGFTNELV QNYWSKQK N IKGVKARFVV TDGGITRVYP KEAGENWQEN PETYEDSFYK RSLDNDNYVF TAPYFNKSGP GAYESGIMVS KAVEIYIQG KLLKPAVVGI KIDVNSWIEN FTKTSIRDPC AGPVCDCKRN SDVMDCVILD DGGFLLMANH DDYTNQIGRF FGEIDPSLMR HLVNISVYA FNKSYDYQSV CEPGAAPKQG AGHRSAYVPS VADILQIGWW ATAAAWSILQ QFLLSLTFPR LLEAVEMEDD D FTASLSKQ SCITEQTQYF FDNDSKSFSG VLDCGNCSRI FHGEKLMNTN LIFIMVESKG TCPCDTRLLI QAEQTSDGPN PC DMVKQPR YRKGPDVCFD NNVLEDYTDC GGVSGLNPSL WYIIGIQFLL LWLVSGSTHR LL |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 4 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 68.3 e/Å2 |
 Movie
Movie Controller
Controller








 Z
Z Y
Y X
X