[English] 日本語
 Yorodumi
Yorodumi- EMDB-34670: The relaxed pre-Tet-S1 state of G264A mutated Tetrahymena group I... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 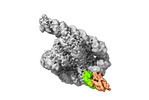 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The relaxed pre-Tet-S1 state of G264A mutated Tetrahymena group I intron with 6nt 3'/5'-exon and 2-aminopurine nucleoside | ||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
| Biological species |   Tetrahymena thermophila (eukaryote) Tetrahymena thermophila (eukaryote) | ||||||||||||||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.73 Å cryo EM / Resolution: 3.73 Å | ||||||||||||||||||||||||
 Authors Authors | Luo B / Zhang C / Ling X / Mukherjee S / Jia G / Xie J / Jia X / Liu L / Baulin EF / Luo Y ...Luo B / Zhang C / Ling X / Mukherjee S / Jia G / Xie J / Jia X / Liu L / Baulin EF / Luo Y / Jiang L / Dong H / Wei X / Bujnicki JM / Su Z | ||||||||||||||||||||||||
| Funding support |  China, China,  Poland, European Union, 7 items Poland, European Union, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Catal / Year: 2023 Journal: Nat Catal / Year: 2023Title: Cryo-EM reveals dynamics of Tetrahymena group I intron self-splicing Authors: Luo B / Zhang C / Ling X / Mukherjee S / Jia G / Xie J / Jia X / Liu L / Baulin EF / Luo Y / Jiang L / Dong H / Wei X / Bujnicki JM / Su Z | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34670.map.gz emd_34670.map.gz | 28.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34670-v30.xml emd-34670-v30.xml emd-34670.xml emd-34670.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34670.png emd_34670.png | 46.9 KB | ||
| Others |  emd_34670_half_map_1.map.gz emd_34670_half_map_1.map.gz emd_34670_half_map_2.map.gz emd_34670_half_map_2.map.gz | 33.1 MB 33.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34670 http://ftp.pdbj.org/pub/emdb/structures/EMD-34670 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34670 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34670 | HTTPS FTP |
-Related structure data
| Related structure data |  8hd6MC  7xd3C  7xd4C  7xd5C  7xd6C  7xd7C  8hd7C  8i7nC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34670.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34670.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_34670_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
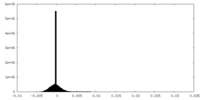
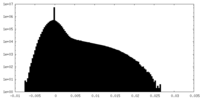
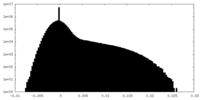
| Density Histograms |
-Half map: #2
| File | emd_34670_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Co-transcriptional folded G264A mutant Tetrahymena group I intron...
| Entire | Name: Co-transcriptional folded G264A mutant Tetrahymena group I intron with 6nt 5'/3'-exon and 2-aminopurine nucleoside |
|---|---|
| Components |
|
-Supramolecule #1: Co-transcriptional folded G264A mutant Tetrahymena group I intron...
| Supramolecule | Name: Co-transcriptional folded G264A mutant Tetrahymena group I intron with 6nt 5'/3'-exon and 2-aminopurine nucleoside type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Tetrahymena thermophila (eukaryote) Tetrahymena thermophila (eukaryote) |
-Macromolecule #1: The relaxed pre-Tet-S1 state molecule of co-transcriptional folde...
| Macromolecule | Name: The relaxed pre-Tet-S1 state molecule of co-transcriptional folded G264A mutant Tetrahymena group I intron with 6nt 3'/5'-exon and 2-aminopurine nucleoside type: rna / ID: 1 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:   Tetrahymena thermophila (eukaryote) Tetrahymena thermophila (eukaryote) |
| Molecular weight | Theoretical: 135.155719 KDa |
| Sequence | String: CUCUCUAAAU AGCAAUAUUU ACCUUUGGAG GGAAAAGUUA UCAGGCAUGC ACCUGGUAGC UAGUCUUUAA ACCAAUAGAU UGCAUCGGU UUAAAAGGCA AGACCGUCAA AUUGCGGGAA AGGGGUCAAC AGCCGUUCAG UACCAAGUCU CAGGGGAAAC U UUGAGAUG ...String: CUCUCUAAAU AGCAAUAUUU ACCUUUGGAG GGAAAAGUUA UCAGGCAUGC ACCUGGUAGC UAGUCUUUAA ACCAAUAGAU UGCAUCGGU UUAAAAGGCA AGACCGUCAA AUUGCGGGAA AGGGGUCAAC AGCCGUUCAG UACCAAGUCU CAGGGGAAAC U UUGAGAUG GCCUUGCAAA GGGUAUGGUA AUAAGCUGAC GGACAUGGUC CUAACCACGC AGCCAAGUCC UAAGUCAACA GA UCUUCUG UUGAUAUGGA UGCAGUUCAC AAACUAAAUG UCGGUCGGGG AAGAUGUAUU CUUCUCAUAA GAUAUAGUCG GAC CUCUCC UUAAUGGGAG CUAGCGGAUG AAGUGAUGCA ACACUGGAGC CGCUGGGAAC UAAUUUGUAU GCGAAAGUAU AUUG AUUAG UUUUGGAGUA CUCG |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 24 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: SPERMIDINE
| Macromolecule | Name: SPERMIDINE / type: ligand / ID: 3 / Number of copies: 1 / Formula: SPD |
|---|---|
| Molecular weight | Theoretical: 145.246 Da |
| Chemical component information |  ChemComp-SPD: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.36 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.7 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 165000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.7 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 165000 |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Temperature | Min: 79.3 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Frames/image: 1-30 / Average electron dose: 55.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 4012136 |
|---|---|
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
| Final 3D classification | Number classes: 6 / Software - Name: RELION (ver. 3.1) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: C1 (asymmetric) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.73 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number images used: 194382 |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8hd6: |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X