+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human RAD52 protein | ||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationdouble-strand break repair via single-strand annealing / DNA double-strand break processing involved in repair via single-strand annealing / DNA recombinase assembly /  mitotic recombination / regulation of nucleotide-excision repair / HDR through MMEJ (alt-NHEJ) / HDR through Single Strand Annealing (SSA) / SUMOylation of DNA damage response and repair proteins / mitotic recombination / regulation of nucleotide-excision repair / HDR through MMEJ (alt-NHEJ) / HDR through Single Strand Annealing (SSA) / SUMOylation of DNA damage response and repair proteins /  protein-DNA complex / double-strand break repair via homologous recombination ...double-strand break repair via single-strand annealing / DNA double-strand break processing involved in repair via single-strand annealing / DNA recombinase assembly / protein-DNA complex / double-strand break repair via homologous recombination ...double-strand break repair via single-strand annealing / DNA double-strand break processing involved in repair via single-strand annealing / DNA recombinase assembly /  mitotic recombination / regulation of nucleotide-excision repair / HDR through MMEJ (alt-NHEJ) / HDR through Single Strand Annealing (SSA) / SUMOylation of DNA damage response and repair proteins / mitotic recombination / regulation of nucleotide-excision repair / HDR through MMEJ (alt-NHEJ) / HDR through Single Strand Annealing (SSA) / SUMOylation of DNA damage response and repair proteins /  protein-DNA complex / double-strand break repair via homologous recombination / double-strand break repair / protein-DNA complex / double-strand break repair via homologous recombination / double-strand break repair /  single-stranded DNA binding / cellular response to oxidative stress / DNA recombination / protein-containing complex / single-stranded DNA binding / cellular response to oxidative stress / DNA recombination / protein-containing complex /  DNA binding / DNA binding /  nucleoplasm / identical protein binding / nucleoplasm / identical protein binding /  nucleus nucleusSimilarity search - Function | ||||||||||||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||||||||
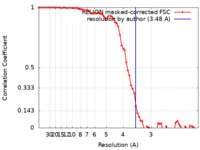
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.48 Å cryo EM / Resolution: 3.48 Å | ||||||||||||||||||||||||
 Authors Authors | Kinoshita C / Takizawa Y / Saotome M / Ogino S / Kurumizaka H / Kagawa W | ||||||||||||||||||||||||
| Funding support |  Japan, 7 items Japan, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: FEBS Open Bio / Year: 2023 Journal: FEBS Open Bio / Year: 2023Title: The cryo-EM structure of full-length RAD52 protein contains an undecameric ring. Authors: Chiaki Kinoshita / Yoshimasa Takizawa / Mika Saotome / Shun Ogino / Hitoshi Kurumizaka / Wataru Kagawa /  Abstract: The human RAD52 protein, which forms an oligomeric ring structure, is involved in DNA double-strand break repair. The N-terminal half of RAD52 is primarily responsible for self-oligomerisation and ...The human RAD52 protein, which forms an oligomeric ring structure, is involved in DNA double-strand break repair. The N-terminal half of RAD52 is primarily responsible for self-oligomerisation and DNA binding. Crystallographic studies have revealed the detailed structure of the N-terminal half. However, only low-resolution structures have been reported for the full-length protein, and thus the structural role of the C-terminal half in self-oligomerisation has remained elusive. In this study, we determined the solution structure of the human RAD52 protein by cryo-electron microscopy (cryo-EM), at an average resolution of 3.5 Å. The structure revealed an undecameric ring that is nearly identical to the crystal structures of the N-terminal half. The cryo-EM map for the C-terminal half was poorly defined, indicating that the region is intrinsically disordered. The present cryo-EM structure provides important insights into the mechanistic roles played by the N-terminal and C-terminal halves of RAD52 during DNA double-strand break repair. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34430.map.gz emd_34430.map.gz | 2.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34430-v30.xml emd-34430-v30.xml emd-34430.xml emd-34430.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34430_fsc.xml emd_34430_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_34430.png emd_34430.png | 231.2 KB | ||
| Others |  emd_34430_half_map_1.map.gz emd_34430_half_map_1.map.gz emd_34430_half_map_2.map.gz emd_34430_half_map_2.map.gz | 23.5 MB 23.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34430 http://ftp.pdbj.org/pub/emdb/structures/EMD-34430 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34430 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34430 | HTTPS FTP |
-Related structure data
| Related structure data |  8h1pMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34430.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34430.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_34430_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
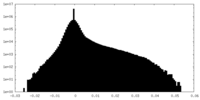
| Density Histograms |
-Half map: #2
| File | emd_34430_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : RAD52
| Entire | Name: RAD52 |
|---|---|
| Components |
|
-Supramolecule #1: RAD52
| Supramolecule | Name: RAD52 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: DNA repair protein RAD52 homolog
| Macromolecule | Name: DNA repair protein RAD52 homolog / type: protein_or_peptide / ID: 1 / Number of copies: 11 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 46.514934 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: GSHMSGTEEA ILGGRDSHPA AGGGSVLCFG QCQYTAEEYQ AIQKALRQRL GPEYISSRMA GGGQKVCYIE GHRVINLANE MFGYNGWAH SITQQNVDFV DLNNGKFYVG VCAFVRVQLK DGSYHEDVGY GVSEGLKSKA LSLEKARKEA VTDGLKRALR S FGNALGNC ...String: GSHMSGTEEA ILGGRDSHPA AGGGSVLCFG QCQYTAEEYQ AIQKALRQRL GPEYISSRMA GGGQKVCYIE GHRVINLANE MFGYNGWAH SITQQNVDFV DLNNGKFYVG VCAFVRVQLK DGSYHEDVGY GVSEGLKSKA LSLEKARKEA VTDGLKRALR S FGNALGNC ILDKDYLRSL NKLPRQLPLE VDLTKAKRQD LEPSVEEARY NSCRPNMALG HPQLQQVTSP SRPSHAVIPA DQ DCSSRSL SSSAVESEAT HQRKLRQKQL QQQFRERMEK QQVRVSTPSA EKSEAAPPAP PVTHSTPVTV SEPLLEKDFL AGV TQELIK TLEDNSEKWA VTPDAGDGVV KPSSRADPAQ TSDTLALNNQ MVTQNRTPHS VCHQKPQAKS GSWDLQTYSA DQRT TGNWE SHRKSQDMKK RKYDPS |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 57.6 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X