+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of human Pannexin isoform 3 | |||||||||
 Map data Map data | Main Map of Pannexin 3 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Pannexins / large pore ion-channels /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationwide pore channel activity / positive regulation of interleukin-1 production /  gap junction / monoatomic cation transport / monoatomic ion transmembrane transport / cell-cell signaling / structural molecule activity / gap junction / monoatomic cation transport / monoatomic ion transmembrane transport / cell-cell signaling / structural molecule activity /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
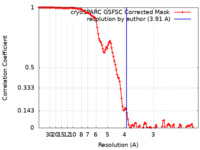
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.91 Å cryo EM / Resolution: 3.91 Å | |||||||||
 Authors Authors | Hussain N / Penmatsa A | |||||||||
| Funding support |  India, 1 items India, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural insights into pore dynamics of human Pannexin isoforms Authors: Hussain N / Apotikar A / Pidathala S / Mukherjee S / Burada AP / Sikdar SK / Vinothkumar KR / Penmatsa A | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34265.map.gz emd_34265.map.gz | 168.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34265-v30.xml emd-34265-v30.xml emd-34265.xml emd-34265.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34265_fsc.xml emd_34265_fsc.xml | 13.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_34265.png emd_34265.png | 64.9 KB | ||
| Filedesc metadata |  emd-34265.cif.gz emd-34265.cif.gz | 6.6 KB | ||
| Others |  emd_34265_half_map_1.map.gz emd_34265_half_map_1.map.gz emd_34265_half_map_2.map.gz emd_34265_half_map_2.map.gz | 165.2 MB 165.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34265 http://ftp.pdbj.org/pub/emdb/structures/EMD-34265 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34265 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34265 | HTTPS FTP |
-Related structure data
| Related structure data |  8gtrMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34265.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34265.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main Map of Pannexin 3 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.065 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Panx3 halfmapB
| File | emd_34265_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Panx3_halfmapB | ||||||||||||
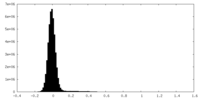
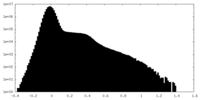
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Panx3 halfmapA
| File | emd_34265_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Panx3_halfmapA | ||||||||||||
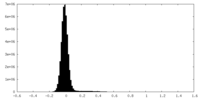
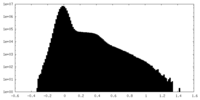
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Pannexin 3
| Entire | Name: Pannexin 3 |
|---|---|
| Components |
|
-Supramolecule #1: Pannexin 3
| Supramolecule | Name: Pannexin 3 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: human isoform 3 of Pannexin. Expressed in ER and Plasma membranes involved in calcium and ATP regulation |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.68 kDa/nm |
-Macromolecule #1: Pannexin-3
| Macromolecule | Name: Pannexin-3 / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.734219 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSLAHTAAEY MLSDALLPDR RGPRLKGLRL ELPLDRIVKF VAVGSPLLLM SLAFAQEFSS GSPISCFSPS NFSIRQAAYV DSSCWDSLL HHKQDGPGQD KMKSLWPHKA LPYSLLALAL LMYLPVLLWQ YAAVPALSSD LLFIISELDK SYNRSIRLVQ H MLKIRQKS ...String: MSLAHTAAEY MLSDALLPDR RGPRLKGLRL ELPLDRIVKF VAVGSPLLLM SLAFAQEFSS GSPISCFSPS NFSIRQAAYV DSSCWDSLL HHKQDGPGQD KMKSLWPHKA LPYSLLALAL LMYLPVLLWQ YAAVPALSSD LLFIISELDK SYNRSIRLVQ H MLKIRQKS SDPYVFWNEL EKARKERYFE FPLLERYLAC KQRSHSLVAT YLLRNSLLLI FTSATYLYLG HFHLDVFFQE EF SCSIKTG LLSDETHVPN LITCRLTSLS IFQIVSLSSV AIYTILVPVI IYNLTRLCRW DKRLLSVYEM LPAFDLLSRK MLG CPINDL NVILLFLRAN ISELISFSWL SVLCVLKDTT TQKHNIDTVV DFMTLLAGLE PSKPKHLTNS ACDEHP UniProtKB:  Pannexin-3 Pannexin-3 |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 7 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #3: PHOSPHATIDYLETHANOLAMINE
| Macromolecule | Name: PHOSPHATIDYLETHANOLAMINE / type: ligand / ID: 3 / Number of copies: 7 / Formula: PTY |
|---|---|
| Molecular weight | Theoretical: 734.039 Da |
| Chemical component information |  ChemComp-PTY: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Fresh solution containing detergent was prepared for every prep. | |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 288 K / Instrument: FEI VITROBOT MARK IV / Details: blotted for 3.5 seconds. | |||||||||||||||
| Details | Sample is homoheptamer purified to homogeneity. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 3.3000000000000003 µm / Nominal defocus min: 1.8 µm / Nominal magnification: 130000 |
| Specialist optics | Phase plate: OTHER / Spherical aberration corrector: None / Chromatic aberration corrector: None / Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV / Details: Bioquantum with K2 camera |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Temperature | Min: 100.0 K / Max: 100.0 K |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-40 / Number grids imaged: 1 / Number real images: 1122 / Average electron dose: 48.8 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 143.6 / Target criteria: Correlation coeficient |
|---|---|
| Output model |  PDB-8gtr: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)