+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GPR110/Gq complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationenergy reserve metabolic process / fat cell differentiation / regulation of lipid metabolic process /  synapse assembly / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor ...energy reserve metabolic process / fat cell differentiation / regulation of lipid metabolic process / synapse assembly / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor ...energy reserve metabolic process / fat cell differentiation / regulation of lipid metabolic process /  synapse assembly / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / synapse assembly / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK /  memory / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / adenylate cyclase-activating dopamine receptor signaling pathway / ADP signalling through P2Y purinoceptor 1 / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / sensory perception of taste / GPER1 signaling / G-protein beta-subunit binding / Inactivation, recovery and regulation of the phototransduction cascade / memory / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / adenylate cyclase-activating dopamine receptor signaling pathway / ADP signalling through P2Y purinoceptor 1 / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / sensory perception of taste / GPER1 signaling / G-protein beta-subunit binding / Inactivation, recovery and regulation of the phototransduction cascade /  heterotrimeric G-protein complex / neuron projection development / G alpha (12/13) signalling events / heterotrimeric G-protein complex / neuron projection development / G alpha (12/13) signalling events /  extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye /  GTPase binding / Ca2+ pathway / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / cytoplasmic vesicle / G alpha (q) signalling events / Ras protein signal transduction / cell population proliferation / Extra-nuclear estrogen signaling / cell surface receptor signaling pathway / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase binding / Ca2+ pathway / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / cytoplasmic vesicle / G alpha (q) signalling events / Ras protein signal transduction / cell population proliferation / Extra-nuclear estrogen signaling / cell surface receptor signaling pathway / G protein-coupled receptor signaling pathway / lysosomal membrane /  GTPase activity / GTPase activity /  synapse / protein-containing complex binding / synapse / protein-containing complex binding /  signal transduction / extracellular exosome / extracellular region / signal transduction / extracellular exosome / extracellular region /  membrane / membrane /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) / Homo sapiens (human) /   Lama glama (llama) Lama glama (llama) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.85 Å cryo EM / Resolution: 2.85 Å | |||||||||
 Authors Authors | He Y / Zhu X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural basis of adhesion GPCR GPR110 activation by stalk peptide and G-proteins coupling. Authors: Xinyan Zhu / Yu Qian / Xiaowan Li / Zhenmei Xu / Ruixue Xia / Na Wang / Jiale Liang / Han Yin / Anqi Zhang / Changyou Guo / Guangfu Wang / Yuanzheng He /  Abstract: Adhesion G protein-coupled receptors (aGPCRs) are keys of many physiological events and attractive targets for various diseases. aGPCRs are also known to be capable of self-activation via an ...Adhesion G protein-coupled receptors (aGPCRs) are keys of many physiological events and attractive targets for various diseases. aGPCRs are also known to be capable of self-activation via an autoproteolysis process that removes the inhibitory GAIN domain on the extracellular side of receptor and releases a stalk peptide to bind and activate the transmembrane side of receptor. However, the detailed mechanism of aGPCR activation remains elusive. Here, we report the cryo-electron microscopy structures of GPR110 (ADGRF1), a member of aGPCR, in complex with G, G, G, G and G The structures reveal distinctive ligand engaging model and activation conformations of GPR110. The structures also unveil the rarely explored GPCR/G and GPCR/G engagements. A comparison of G, G, G, G and G engagements with GPR110 reveals details of G-protein engagement, including a dividing point at the far end of the alpha helix 5 (αH5) of Gα subunit that separates G/G engagements from G/G/G engagements. This is also where G/G bind the receptor through both hydrophobic and polar interaction, while G/G/G engage receptor mainly through hydrophobic interaction. We further provide physiological evidence of GPR110 activation via stalk peptide. Taken together, our study fills the missing information of GPCR/G-protein engagement and provides a framework for understanding aGPCR activation and GPR110 signaling. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32881.map.gz emd_32881.map.gz | 55.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32881-v30.xml emd-32881-v30.xml emd-32881.xml emd-32881.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
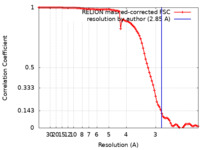
| FSC (resolution estimation) |  emd_32881_fsc.xml emd_32881_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_32881.png emd_32881.png | 74.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32881 http://ftp.pdbj.org/pub/emdb/structures/EMD-32881 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32881 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32881 | HTTPS FTP |
-Related structure data
| Related structure data |  7wxuMC  7wxwC  7wy0C  7wz7C  7x2vC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32881.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32881.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : GPCR/G-protein complex
| Entire | Name: GPCR/G-protein complex |
|---|---|
| Components |
|
-Supramolecule #1: GPCR/G-protein complex
| Supramolecule | Name: GPCR/G-protein complex / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #2: GPCR/G-protein
| Supramolecule | Name: GPCR/G-protein / type: complex / Chimera: Yes / ID: 2 / Parent: 1 / Macromolecule list: #1-#3, #5 |
|---|---|
| Source (natural) | Organism:   Lama glama (llama) Lama glama (llama) |
| Recombinant expression | Organism:  Bacteria abnormis (insect) Bacteria abnormis (insect) |
-Supramolecule #3: NB35
| Supramolecule | Name: NB35 / type: complex / Chimera: Yes / ID: 3 / Parent: 1 / Macromolecule list: #4 |
|---|
-Macromolecule #1: engineered mini Galpha-Q subunit
| Macromolecule | Name: engineered mini Galpha-Q subunit / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.855578 KDa |
| Recombinant expression | Organism:  Insect BA phytoplasma (bacteria) Insect BA phytoplasma (bacteria) |
| Sequence | String: MMGCTLSAED KAAVERSKMI EKQLQKDKQV YRRTLRLLLL GADNSGKSTI VKQMRIYHVN GYSEEECKQY KAVVYSNTIQ SIIAIIRAM GRLKIDFGDS ARADDARQLF VLAGAAEEGF MTAELAGVIK RLWKDSGVQA CFNRSREYQL NDSAAYYLND L DRIAQPNY ...String: MMGCTLSAED KAAVERSKMI EKQLQKDKQV YRRTLRLLLL GADNSGKSTI VKQMRIYHVN GYSEEECKQY KAVVYSNTIQ SIIAIIRAM GRLKIDFGDS ARADDARQLF VLAGAAEEGF MTAELAGVIK RLWKDSGVQA CFNRSREYQL NDSAAYYLND L DRIAQPNY IPTQQDVLRT RVKTSGIFET KFQVDKVNFH MFDVGAQRDE RRKWIQCFND VTAIIFVVDS SDYNRLQEAL ND FKSIWNN RWLRTISVIL FLNKQDLLAE KVLAGKSKIE DYFPEFARYT TPEDATPEPG EDPRVTRAKY FIRKEFVDIS TAS GDGRHI CYPHFTCSVD TENARRIFND CKDIILQMNL REYNLV |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.915496 KDa |
| Recombinant expression | Organism:  Insect BA phytoplasma (bacteria) Insect BA phytoplasma (bacteria) |
| Sequence | String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWN |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Insect BA phytoplasma (bacteria) Insect BA phytoplasma (bacteria) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L |
-Macromolecule #4: NB35
| Macromolecule | Name: NB35 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Lama glama (llama) Lama glama (llama) |
| Molecular weight | Theoretical: 17.381584 KDa |
| Recombinant expression | Organism:  Bacteria abnormis (insect) Bacteria abnormis (insect) |
| Sequence | String: MKYLLPTAAA GLLLLAAQPA MAQVQLQESG GGLVQPGGSL RLSCAASGFT FSNYKMNWVR QAPGKGLEWV SDISQSGASI SYTGSVKGR FTISRDNAKN TLYLQMNSLK PEDTAVYYCA RCPAPFTRDC FDVTSTTYAY RGQGTQVTVS SAAALEHHHH H H |
-Macromolecule #5: Adhesion G-protein coupled receptor F1
| Macromolecule | Name: Adhesion G-protein coupled receptor F1 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 101.465219 KDa |
| Recombinant expression | Organism:  Insect BA phytoplasma (bacteria) Insect BA phytoplasma (bacteria) |
| Sequence | String: MKVGVLWLIS FFTFTDGHGG FLGKNDGIKT KKELIVNKKK HLGPVEEYQL LLQVTYRDSK EKRDLRNFLK LLKPPLLWSH GLIRIIRAK ATTDCNSLNG VLQCTCEDSY TWFPPSCLDP QNCYLHTAGA LPSCECHLNN LSQSVNFCER TKIWGTFKIN E RFTNDLLN ...String: MKVGVLWLIS FFTFTDGHGG FLGKNDGIKT KKELIVNKKK HLGPVEEYQL LLQVTYRDSK EKRDLRNFLK LLKPPLLWSH GLIRIIRAK ATTDCNSLNG VLQCTCEDSY TWFPPSCLDP QNCYLHTAGA LPSCECHLNN LSQSVNFCER TKIWGTFKIN E RFTNDLLN SSSAIYSKYA NGIEIQLKKA YERIQGFESV QVTQFRNGSI VAGYEVVGSS SASELLSAIE HVAEKAKTAL HK LFPLEDG SFRVFGKAQC NDIVFGFGSK DDEYTLPCSS GYRGNITAKC ESSGWQVIRE TCVLSLLEEL NKNFSMIVGN ATE AAVSSF VQNLSVIIRQ NPSTTVGNLA SVVSILSNIS SLSLASHFRV SNSTMEDVIS IADNILNSAS VTNWTVLLRE EKYA SSRLL ETLENISTLV PPTALPLNFS RKFIDWKGIP VNKSQLKRGY SYQIKMCPQN TSIPIRGRVL IGSDQFQRSL PETII SMAS LTLGNILPVS KNGNAQVNGP VISTVIQNYS INEVFLFFSK IESNLSQPHC VFWDFSHLQW NDAGCHLVNE TQDIVT CQC THLTSFSILM SPFVPSTIFP VVKWITYVGL GISIGSLILC LIIEALFWKQ IKKSQTSHTR RICMVNIALS LLIADVW FI VGATVDTTVN PSGVCTAAVF FTHFFYLSLF FWMLMLGILL AYRIILVFHH MAQHLMMAVG FCLGYGCPLI ISVITIAV T QPSNTYKRKD VCWLNWSNGS KPLLAFVVPA LAIVAVNFVV VLLVLTKLWR PTVGERLSRD DKATIIRVGK SLLILTPLL GLTWGFGIGT IVDSQNLAWH VIFALLNAFQ GFFILCFGIL LDSKLRQLLF NKLSALSSWK QTEKQNSSDL SAKPKFSKPF NPLQNKGHY AFSHTGDSSD NIMLTQFVSN E |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 1.2 µm Bright-field microscopy / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 1.2 µm |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller