[English] 日本語
 Yorodumi
Yorodumi- EMDB-28993: Cryo-EM structure of Cascade-DNA-TniQ-TnsC complex in type I-B CA... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
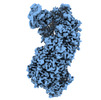
| Title | Cryo-EM structure of Cascade-DNA-TniQ-TnsC complex in type I-B CAST system | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  CRISPR / CRISPR /  DNA recognition / DNA recognition /  DNA BINDING PROTEIN DNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |  Peltigera membranacea (fungus) / Peltigera membranacea (fungus) /  Nostoc sp. 'Peltigera membranacea cyanobiont' 210A (bacteria) / synthetic construct (others) Nostoc sp. 'Peltigera membranacea cyanobiont' 210A (bacteria) / synthetic construct (others) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.19 Å cryo EM / Resolution: 3.19 Å | |||||||||
 Authors Authors | Chang L / Wang S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Molecular mechanism for Tn7-like transposon recruitment by a type I-B CRISPR effector. Authors: Shukun Wang / Clinton Gabel / Romana Siddique / Thomas Klose / Leifu Chang /  Abstract: Tn7-like transposons have co-opted CRISPR-Cas systems to facilitate the movement of their own DNA. These CRISPR-associated transposons (CASTs) are promising tools for programmable gene knockin. A key ...Tn7-like transposons have co-opted CRISPR-Cas systems to facilitate the movement of their own DNA. These CRISPR-associated transposons (CASTs) are promising tools for programmable gene knockin. A key feature of CASTs is their ability to recruit Tn7-like transposons to nuclease-deficient CRISPR effectors. However, how Tn7-like transposons are recruited by diverse CRISPR effectors remains poorly understood. Here, we present the cryo-EM structure of a recruitment complex comprising the Cascade complex, TniQ, TnsC, and the target DNA in the type I-B CAST from Peltigera membranacea cyanobiont 210A. Target DNA recognition by Cascade induces conformational changes in Cas6 and primes TniQ recruitment through its C-terminal domain. The N-terminal domain of TniQ is bound to the seam region of the TnsC spiral heptamer. Our findings provide insights into the diverse mechanisms for the recruitment of Tn7-like transposons to CRISPR effectors and will aid in the development of CASTs as gene knockin tools. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28993.map.gz emd_28993.map.gz | 16.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28993-v30.xml emd-28993-v30.xml emd-28993.xml emd-28993.xml | 24.8 KB 24.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28993.png emd_28993.png | 134.6 KB | ||
| Filedesc metadata |  emd-28993.cif.gz emd-28993.cif.gz | 7.3 KB | ||
| Others |  emd_28993_half_map_1.map.gz emd_28993_half_map_1.map.gz emd_28993_half_map_2.map.gz emd_28993_half_map_2.map.gz | 200.2 MB 200.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28993 http://ftp.pdbj.org/pub/emdb/structures/EMD-28993 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28993 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28993 | HTTPS FTP |
-Related structure data
| Related structure data |  8fcuMC  8fcjC  8fcvC  8fcwC  8fcxC  8fd2C  8fd3C  8ff4C  8ff5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28993.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28993.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.054 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_28993_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_28993_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Cascade-DNA-TniQ-TnsC complex
+Supramolecule #1: Cascade-DNA-TniQ-TnsC complex
+Macromolecule #1: Type I-B CRISPR-associated protein Cas5
+Macromolecule #2: Type I-B CRISPR-associated protein Cas6
+Macromolecule #3: Type I-B CRISPR-associated protein Cas7
+Macromolecule #4: Type I-MYXAN CRISPR-associated Cas8a1/Cmx1
+Macromolecule #5: Type I-B CRISPR-associated protein Cas11
+Macromolecule #9: TniQ
+Macromolecule #10: TnsC
+Macromolecule #6: RNA
+Macromolecule #7: Target DNA strand
+Macromolecule #8: Non-target DNA strand
+Macromolecule #11: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #12: MAGNESIUM ION
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 54.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.19 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 96534 |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X