+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
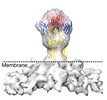
| Title | Cryo-EM map of Chikungunya virus strain s27 | |||||||||
 Map data Map data | Chikungunya virus strain S27 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Chikungunya virus strain S27-African prototype Chikungunya virus strain S27-African prototype | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 6.75 Å cryo EM / Resolution: 6.75 Å | |||||||||
 Authors Authors | Mangala Prasad V / Lee KK | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Visualization of conformational changes and membrane remodeling leading to genome delivery by viral class-II fusion machinery. Authors: Vidya Mangala Prasad / Jelle S Blijleven / Jolanda M Smit / Kelly K Lee /    Abstract: Chikungunya virus (CHIKV) is a human pathogen that delivers its genome to the host cell cytoplasm through endocytic low pH-activated membrane fusion mediated by class-II fusion proteins. Though ...Chikungunya virus (CHIKV) is a human pathogen that delivers its genome to the host cell cytoplasm through endocytic low pH-activated membrane fusion mediated by class-II fusion proteins. Though structures of prefusion, icosahedral CHIKV are available, structural characterization of virion interaction with membranes has been limited. Here, we have used cryo-electron tomography to visualize CHIKV's complete membrane fusion pathway, identifying key intermediary glycoprotein conformations coupled to membrane remodeling events. Using sub-tomogram averaging, we elucidate features of the low pH-exposed virion, nucleocapsid and full-length E1-glycoprotein's post-fusion structure. Contrary to class-I fusion systems, CHIKV achieves membrane apposition by protrusion of extended E1-glycoprotein homotrimers into the target membrane. The fusion process also features a large hemifusion diaphragm that transitions to a wide pore for intact nucleocapsid delivery. Our analyses provide comprehensive ultrastructural insights into the class-II virus fusion system function and direct mechanistic characterization of the fundamental process of protein-mediated membrane fusion. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27559.map.gz emd_27559.map.gz | 189.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27559-v30.xml emd-27559-v30.xml emd-27559.xml emd-27559.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27559_fsc.xml emd_27559_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_27559.png emd_27559.png | 222.8 KB | ||
| Others |  emd_27559_half_map_1.map.gz emd_27559_half_map_1.map.gz emd_27559_half_map_2.map.gz emd_27559_half_map_2.map.gz | 191 MB 190.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27559 http://ftp.pdbj.org/pub/emdb/structures/EMD-27559 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27559 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27559 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27559.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27559.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Chikungunya virus strain S27 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 2.7 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half-map
| File | emd_27559_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map
| File | emd_27559_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Chikungunya virus strain S27-African prototype
| Entire | Name:   Chikungunya virus strain S27-African prototype Chikungunya virus strain S27-African prototype |
|---|---|
| Components |
|
-Supramolecule #1: Chikungunya virus strain S27-African prototype
| Supramolecule | Name: Chikungunya virus strain S27-African prototype / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 371094 Sci species name: Chikungunya virus strain S27-African prototype Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:   Aedes aegypti (yellow fever mosquito) Aedes aegypti (yellow fever mosquito) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Component - Concentration: 1.0 X / Component - Formula: HBS / Component - Name: Hepes Buffer Saline Details: Hepes Buffer Saline (10mM HEPES, 150mM NaCl, 50mM sodium citrate) |
|---|---|
| Grid | Model: EMS Lacey Carbon / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: LACEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 105000 Bright-field microscopy / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 105000 |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Number real images: 495 / Average exposure time: 0.2 sec. / Average electron dose: 43.9 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X




















