+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cereblon~DDB1 bound to Pomalidomide | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of monoatomic ion transmembrane transport / positive regulation by virus of viral protein levels in host cell / epigenetic programming in the zygotic pronuclei / spindle assembly involved in female meiosis / Cul4-RING E3 ubiquitin ligase complex / UV-damage excision repair / biological process involved in interaction with symbiont / regulation of mitotic cell cycle phase transition / WD40-repeat domain binding / Cul4A-RING E3 ubiquitin ligase complex ...negative regulation of monoatomic ion transmembrane transport / positive regulation by virus of viral protein levels in host cell / epigenetic programming in the zygotic pronuclei / spindle assembly involved in female meiosis / Cul4-RING E3 ubiquitin ligase complex / UV-damage excision repair / biological process involved in interaction with symbiont / regulation of mitotic cell cycle phase transition / WD40-repeat domain binding / Cul4A-RING E3 ubiquitin ligase complex / Cul4B-RING E3 ubiquitin ligase complex / ubiquitin ligase complex scaffold activity / locomotory exploration behavior / negative regulation of reproductive process / negative regulation of developmental process / cullin family protein binding / positive regulation of Wnt signaling pathway / viral release from host cell / ectopic germ cell programmed cell death / positive regulation of viral genome replication / negative regulation of protein-containing complex assembly / positive regulation of gluconeogenesis / proteasomal protein catabolic process / Recognition of DNA damage by PCNA-containing replication complex / nucleotide-excision repair / positive regulation of protein-containing complex assembly / DNA Damage Recognition in GG-NER / Dual Incision in GG-NER /  regulation of circadian rhythm / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / Formation of TC-NER Pre-Incision Complex / regulation of circadian rhythm / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / Formation of TC-NER Pre-Incision Complex /  Wnt signaling pathway / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / positive regulation of protein catabolic process / cellular response to UV / rhythmic process / protein-macromolecule adaptor activity / site of double-strand break / Wnt signaling pathway / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / positive regulation of protein catabolic process / cellular response to UV / rhythmic process / protein-macromolecule adaptor activity / site of double-strand break /  Neddylation / ubiquitin-dependent protein catabolic process / proteasome-mediated ubiquitin-dependent protein catabolic process / Potential therapeutics for SARS / transmembrane transporter binding / damaged DNA binding / Neddylation / ubiquitin-dependent protein catabolic process / proteasome-mediated ubiquitin-dependent protein catabolic process / Potential therapeutics for SARS / transmembrane transporter binding / damaged DNA binding /  chromosome, telomeric region / protein ubiquitination / chromosome, telomeric region / protein ubiquitination /  DNA repair / apoptotic process / DNA damage response / protein-containing complex binding / DNA repair / apoptotic process / DNA damage response / protein-containing complex binding /  nucleolus / negative regulation of apoptotic process / perinuclear region of cytoplasm / protein-containing complex / nucleolus / negative regulation of apoptotic process / perinuclear region of cytoplasm / protein-containing complex /  DNA binding / DNA binding /  extracellular space / extracellular exosome / extracellular space / extracellular exosome /  nucleoplasm / nucleoplasm /  membrane / membrane /  metal ion binding / metal ion binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.9 Å cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Watson ER / Lander GC | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Molecular glue CELMoD compounds are regulators of cereblon conformation. Authors: Edmond R Watson / Scott Novick / Mary E Matyskiela / Philip P Chamberlain / Andres H de la Peña / Jinyi Zhu / Eileen Tran / Patrick R Griffin / Ingrid E Wertz / Gabriel C Lander /  Abstract: Cereblon (CRBN) is a ubiquitin ligase (E3) substrate receptor protein co-opted by CRBN E3 ligase modulatory drug (CELMoD) agents that target therapeutically relevant proteins for degradation. Prior ...Cereblon (CRBN) is a ubiquitin ligase (E3) substrate receptor protein co-opted by CRBN E3 ligase modulatory drug (CELMoD) agents that target therapeutically relevant proteins for degradation. Prior crystallographic studies defined the drug-binding site within CRBN's thalidomide-binding domain (TBD), but the allostery of drug-induced neosubstrate binding remains unclear. We performed cryo-electron microscopy analyses of the DNA damage-binding protein 1 (DDB1)-CRBN apo complex and compared these structures with DDB1-CRBN in the presence of CELMoD compounds alone and complexed with neosubstrates. Association of CELMoD compounds to the TBD is necessary and sufficient for triggering CRBN allosteric rearrangement from an open conformation to the canonical closed conformation. The neosubstrate Ikaros only stably associates with the closed CRBN conformation, illustrating the importance of allostery for CELMoD compound efficacy and informing structure-guided design strategies to improve therapeutic efficacy. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27242.map.gz emd_27242.map.gz | 40.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27242-v30.xml emd-27242-v30.xml emd-27242.xml emd-27242.xml | 20.9 KB 20.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27242_fsc.xml emd_27242_fsc.xml | 7.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_27242.png emd_27242.png | 154.5 KB | ||
| Others |  emd_27242_additional_1.map.gz emd_27242_additional_1.map.gz emd_27242_half_map_1.map.gz emd_27242_half_map_1.map.gz emd_27242_half_map_2.map.gz emd_27242_half_map_2.map.gz | 21.2 MB 39.8 MB 39.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27242 http://ftp.pdbj.org/pub/emdb/structures/EMD-27242 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27242 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27242 | HTTPS FTP |
-Related structure data
| Related structure data |  8d81MC  8cvpC  8d7uC  8d7vC  8d7wC  8d7xC  8d7yC  8d7zC  8d80C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27242.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27242.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.91 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_27242_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27242_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_27242_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cereblon~DDB1 bound to Pomalidomide
| Entire | Name: Cereblon~DDB1 bound to Pomalidomide |
|---|---|
| Components |
|
-Supramolecule #1: Cereblon~DDB1 bound to Pomalidomide
| Supramolecule | Name: Cereblon~DDB1 bound to Pomalidomide / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 143 KDa |
-Macromolecule #1: DNA damage-binding protein 1
| Macromolecule | Name: DNA damage-binding protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 93.347078 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MSYNYVVTAQ KPTAVNGCVT GHFTSAEDLN LLIAKNTRLE IYVVTAEGLR PVKEVGMYGK IAVMELFRPK GESKDLLFIL TAKYNACIL EYKQSGESID IITRAHGNVQ DRIGRPSETG IIGIIDPECR MIGLRLYDGL FKVIPLDRDN KELKAFNIRL E ELHVIDVK ...String: MSYNYVVTAQ KPTAVNGCVT GHFTSAEDLN LLIAKNTRLE IYVVTAEGLR PVKEVGMYGK IAVMELFRPK GESKDLLFIL TAKYNACIL EYKQSGESID IITRAHGNVQ DRIGRPSETG IIGIIDPECR MIGLRLYDGL FKVIPLDRDN KELKAFNIRL E ELHVIDVK FLYGCQAPTI CFVYQDPQGR HVKTYEVSLR EKEFNKGPWK QENVEAEASM VIAVPEPFGG AIIIGQESIT YH NGDKYLA IAPPIIKQST IVCHNRVDPN GSRYLLGDME GRLFMLLLEK EEQMDGTVTL KDLRVELLGE TSIAECLTYL DNG VVFVGS RLGDSQLVKL NVDSNEQGSY VVAMETFTNL GPIVDMCVVD LERQGQGQLV TCSGAFKEGS LRIIRNGIGG NGNS GEIQK LHIRTVPLYE SPRKICYQEV SQCFGVLSSR IEVQDTSGGT TALRPSASTQ ALSSSVSSSK LFSSSTAPHE TSFGE EVEV HNLLIIDQHT FEVLHAHQFL QNEYALSLVS CKLGKDPNTY FIVGTAMVYP EEAEPKQGRI VVFQYSDGKL QTVAEK EVK GAVYSMVEFN GKLLASINST VRLYEWTTEK ELRTECNHYN NIMALYLKTK GDFILVGDLM RSVLLLAYKP MEGNFEE IA RDFNPNWMSA VEILDDDNFL GAENAFNLFV CQKDSAATTD EERQHLQEVG LFHLGEFVNV FCHGSLVMQN LGETSTPT Q GSVLFGTVNG MIGLVTSLSE SWYNLLLDMQ NRLNKVIKSV GKIEHSFWRS FHTERKTEPA TGFIDGDLIE SFLDISRPK MQEVVANLQY DDGSGMKREA TADDLIKVVE ELTRIH |
-Macromolecule #2: Protein cereblon
| Macromolecule | Name: Protein cereblon / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.603676 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MAGEGDQQDA AHNMGNHLPL LPAESEEEDE MEVEDQDSKE AKKPNIINFD TSLPTSHTYL GADMEEFHGR TLHDDDSCQV IPVLPQVMM ILIPGQTLPL QLFHPQEVSM VRNLIQKDRT FAVLAYSNVQ EREAQFGTTA EIYAYREEQD FGIEIVKVKA I GRQRFKVL ...String: MAGEGDQQDA AHNMGNHLPL LPAESEEEDE MEVEDQDSKE AKKPNIINFD TSLPTSHTYL GADMEEFHGR TLHDDDSCQV IPVLPQVMM ILIPGQTLPL QLFHPQEVSM VRNLIQKDRT FAVLAYSNVQ EREAQFGTTA EIYAYREEQD FGIEIVKVKA I GRQRFKVL ELRTQSDGIQ QAKVQILPEC VLPSTMSAVQ LESLNKCQIF PSKPVSREDQ CSYKWWQKYQ KRKFHCANLT SW PRWLYSL YDAETLMDRI KKQLREWDEN LKDDSLPSNP IDFSYRVAAC LPIDDVLRIQ LLKIGSAIQR LRCELDIMNK CTS LCCKQC QETEITTKNE IFSLSLCGPM AAYVNPHGYV HETLTVYKAC NLNLIGRPST EHSWFPGYAW TVAQCKICAS HIGW KFTAT KKDMSPQKFW GLTRSALLPT IPDTEDEISP DKVILCL |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: S-Pomalidomide
| Macromolecule | Name: S-Pomalidomide / type: ligand / ID: 4 / Number of copies: 1 / Formula: Y70 |
|---|---|
| Molecular weight | Theoretical: 273.244 Da |
| Chemical component information | 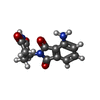 ChemComp-Y70: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 20 mg/mL |
|---|---|
| Buffer | pH: 7 / Details: 20mM HEPES pH 7, 240mM NaCl, 3mM TCEP |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER / Details: Manual plunge in 4 degree C cold room. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 62.5 e/Å2 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z
Z Y
Y X
X


























