+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of NKCC1 Bu-I | |||||||||
 Map data Map data | Cryo-EM structure of NKCC1 Bu-I | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cotransporter / dimer /  ion / ion /  MEMBRANE PROTEIN / MEMBRANE PROTEIN /  Transport Protein Transport Protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of cell volume / positive regulation of aspartate secretion / transepithelial ammonium transport / regulation of matrix metallopeptidase secretion / cell body membrane / inorganic anion import across plasma membrane / inorganic cation import across plasma membrane / chloride:monoatomic cation symporter activity / sodium:potassium:chloride symporter activity / metal ion transmembrane transporter activity ...positive regulation of cell volume / positive regulation of aspartate secretion / transepithelial ammonium transport / regulation of matrix metallopeptidase secretion / cell body membrane / inorganic anion import across plasma membrane / inorganic cation import across plasma membrane / chloride:monoatomic cation symporter activity / sodium:potassium:chloride symporter activity / metal ion transmembrane transporter activity / transepithelial chloride transport / potassium ion transmembrane transporter activity / Cation-coupled Chloride cotransporters / ammonium transmembrane transport / sodium ion homeostasis / intracellular chloride ion homeostasis / negative regulation of vascular wound healing / chloride ion homeostasis / cellular response to potassium ion / ammonium channel activity / intracellular potassium ion homeostasis / cell projection membrane / intracellular sodium ion homeostasis / sodium ion import across plasma membrane / potassium ion homeostasis / cellular response to chemokine / T cell chemotaxis / hyperosmotic response / gamma-aminobutyric acid signaling pathway / cell volume homeostasis / regulation of spontaneous synaptic transmission / maintenance of blood-brain barrier / potassium ion import across plasma membrane / transport across blood-brain barrier / sodium ion transmembrane transport / lateral plasma membrane / monoatomic ion transport / chloride transmembrane transport / basal plasma membrane / cell projection / cell periphery /  Hsp90 protein binding / cytoplasmic vesicle membrane / Hsp90 protein binding / cytoplasmic vesicle membrane /  extracellular vesicle / extracellular vesicle /  cell body / protein-folding chaperone binding / basolateral plasma membrane / neuron projection / apical plasma membrane / neuronal cell body / cell body / protein-folding chaperone binding / basolateral plasma membrane / neuron projection / apical plasma membrane / neuronal cell body /  protein kinase binding / extracellular exosome / protein kinase binding / extracellular exosome /  membrane / membrane /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
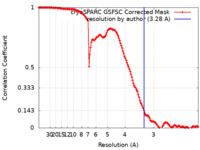
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.28 Å cryo EM / Resolution: 3.28 Å | |||||||||
 Authors Authors | Moseng MA | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Inhibition mechanism of NKCC1 involves the carboxyl terminus and long-range conformational coupling. Authors: Mitchell A Moseng / Chih-Chia Su / Kerri Rios / Meng Cui / Meinan Lyu / Przemyslaw Glaza / Philip A Klenotic / Eric Delpire / Edward W Yu /  Abstract: The Na-K-2Cl cotransporter-1 (NKCC1) is an electroneutral Na-dependent transporter responsible for simultaneously translocating Na, K, and Cl ions into cells. In human tissue, NKCC1 plays a critical ...The Na-K-2Cl cotransporter-1 (NKCC1) is an electroneutral Na-dependent transporter responsible for simultaneously translocating Na, K, and Cl ions into cells. In human tissue, NKCC1 plays a critical role in regulating cytoplasmic volume, fluid intake, chloride homeostasis, and cell polarity. Here, we report four structures of human NKCC1 (hNKCC1), both in the absence and presence of loop diuretic (bumetanide or furosemide), using single-particle cryo-electron microscopy. These structures allow us to directly observe various novel conformations of the hNKCC1 dimer. They also reveal two drug-binding sites located at the transmembrane and cytosolic carboxyl-terminal domains, respectively. Together, our findings enable us to delineate an inhibition mechanism that involves a coupled movement between the cytosolic and transmembrane domains of hNKCC1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25204.map.gz emd_25204.map.gz | 230 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25204-v30.xml emd-25204-v30.xml emd-25204.xml emd-25204.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25204_fsc.xml emd_25204_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_25204.png emd_25204.png | 149.5 KB | ||
| Masks |  emd_25204_msk_1.map emd_25204_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Others |  emd_25204_half_map_1.map.gz emd_25204_half_map_1.map.gz emd_25204_half_map_2.map.gz emd_25204_half_map_2.map.gz | 226.6 MB 226.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25204 http://ftp.pdbj.org/pub/emdb/structures/EMD-25204 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25204 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25204 | HTTPS FTP |
-Related structure data
| Related structure data |  7smpMC  7mxoC  7n3nC  7sflC  8steC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25204.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25204.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of NKCC1 Bu-I | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25204_msk_1.map emd_25204_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 1
| File | emd_25204_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_25204_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : NKCC1 Bu-I
| Entire | Name: NKCC1 Bu-I |
|---|---|
| Components |
|
-Supramolecule #1: NKCC1 Bu-I
| Supramolecule | Name: NKCC1 Bu-I / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Solute carrier family 12 member 2
| Macromolecule | Name: Solute carrier family 12 member 2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 103.313422 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVKFGWIKGV LVRCMLNIWG VMLFIRLSWI VGQAGIGLSV LVIMMATVVT TITGLSTSAI ATNGFVRGGG AYYLISRSLG PEFGGAIGL IFAFANAVAV AMYVVGFAET VVELLKEHSI LMIDEINDIR IIGAITVVIL LGISVAGMEW EAKAQIVLLV I LLLAIGDF ...String: AVKFGWIKGV LVRCMLNIWG VMLFIRLSWI VGQAGIGLSV LVIMMATVVT TITGLSTSAI ATNGFVRGGG AYYLISRSLG PEFGGAIGL IFAFANAVAV AMYVVGFAET VVELLKEHSI LMIDEINDIR IIGAITVVIL LGISVAGMEW EAKAQIVLLV I LLLAIGDF VIGTFIPLES KKPKGFFGYK SEIFNENFGP DFREEETFFS VFAIFFPAAT GILAGANISG DLADPQSAIP KG TLLAILI TTLVYVGIAV SVGSCVVRDA TGNVNDTIVT ELTNCTSAAC KLNFDFSSCE SSPCSYGLMN NFQVMSMVSG FTP LISAGI FSATLSSALA SLVSAPKIFQ ALCKDNIYPA FQMFAKGYGK NNEPLRGYIL TFLIALGFIL IAELNVIAPI ISNF FLASY ALINFSVFHA SLAKSPGWRP AFKYYNMWIS LLGAILCCIV MFVINWWAAL LTYVIVLGLY IYVTYKKPDV NWGSS TQAL TYLNALQHSI RLSGVEDHVK NFRPQCLVMT GAPNSRPALL HLVHDFTKNV GLMICGHVHM GPRRQAMKEM SIDQAK YQR WLIKNKMKAF YAPVHADDLR EGAQYLMQAA GLGRMKPNTL VLGFKKDWLQ ADMRDVDMYI NLFHDAFDIQ YGVVVIR LK EGLDISHLQG QEELLSSQEK SPGTKDVVVS VEYSKKSDLD TSKPLSEKPI THKVEEEDGK TATQPLLKKE SKGPIVPL N VADQKLLEAS TQFQKKQGKN TIDVWWLFDD GGLTLLIPYL LTTKKKWKDC KIRVFIGGKI NRIDHDRRAM ATLLSKFRI DFSDIMVLGD INTKPKKENI IAFEEIIEPY RLHEDDKEQD IADKMKEDEP WRITDNELEL YKTKTYRQIR LNELLKEHSS TANIIVMSL PVARKGAVSS ALYMAWLEAL SKDLPPILLV RGNHQSVLTF Y |
-Macromolecule #2: 3-(butylamino)-4-phenoxy-5-sulfamoylbenzoic acid
| Macromolecule | Name: 3-(butylamino)-4-phenoxy-5-sulfamoylbenzoic acid / type: ligand / ID: 2 / Number of copies: 3 / Formula: 82U |
|---|---|
| Molecular weight | Theoretical: 364.416 Da |
| Chemical component information | 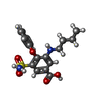 ChemComp-82U: |
-Macromolecule #3: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.2 µm Bright-field microscopy / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.2 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X