[English] 日本語
 Yorodumi
Yorodumi- EMDB-25037: Cryo-EM 3D map of the human Exostosin-1 and Exostosin-2 heterodim... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM 3D map of the human Exostosin-1 and Exostosin-2 heterodimer in complex with a 7-sugar oligosaccharide acceptor analog | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase / glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase /  N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase / N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase /  hypersensitivity / heart field specification / lymphocyte adhesion to endothelial cell of high endothelial venule / heparan sulfate N-acetylglucosaminyltransferase activity / hypersensitivity / heart field specification / lymphocyte adhesion to endothelial cell of high endothelial venule / heparan sulfate N-acetylglucosaminyltransferase activity /  glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase activity / glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase activity /  N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase activity / smoothened signaling pathway involved in lung development / developmental growth involved in morphogenesis ... N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase activity / smoothened signaling pathway involved in lung development / developmental growth involved in morphogenesis ... glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase / glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase /  N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase / N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase /  hypersensitivity / heart field specification / lymphocyte adhesion to endothelial cell of high endothelial venule / heparan sulfate N-acetylglucosaminyltransferase activity / hypersensitivity / heart field specification / lymphocyte adhesion to endothelial cell of high endothelial venule / heparan sulfate N-acetylglucosaminyltransferase activity /  glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase activity / glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase activity /  N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase activity / smoothened signaling pathway involved in lung development / developmental growth involved in morphogenesis / sweat gland development / perichondral bone morphogenesis / mesenchymal cell differentiation involved in bone development / response to leukemia inhibitory factor / chondroitin sulfate metabolic process / UDP-N-acetylglucosamine transferase complex / response to heparin / chondrocyte hypertrophy / embryonic skeletal joint development / hematopoietic stem cell migration to bone marrow / : / heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process / fluid transport / N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase activity / smoothened signaling pathway involved in lung development / developmental growth involved in morphogenesis / sweat gland development / perichondral bone morphogenesis / mesenchymal cell differentiation involved in bone development / response to leukemia inhibitory factor / chondroitin sulfate metabolic process / UDP-N-acetylglucosamine transferase complex / response to heparin / chondrocyte hypertrophy / embryonic skeletal joint development / hematopoietic stem cell migration to bone marrow / : / heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process / fluid transport /  glucuronosyltransferase activity / limb joint morphogenesis / tight junction organization / heparin biosynthetic process / Defective EXT2 causes exostoses 2 / Defective EXT1 causes exostoses 1, TRPS2 and CHDS / stomach development / sebaceous gland development / glomerular basement membrane development / dendrite self-avoidance / heparan sulfate proteoglycan biosynthetic process / HS-GAG biosynthesis / glycosaminoglycan biosynthetic process / lymphocyte migration into lymphoid organs / chondrocyte proliferation / glucuronosyltransferase activity / limb joint morphogenesis / tight junction organization / heparin biosynthetic process / Defective EXT2 causes exostoses 2 / Defective EXT1 causes exostoses 1, TRPS2 and CHDS / stomach development / sebaceous gland development / glomerular basement membrane development / dendrite self-avoidance / heparan sulfate proteoglycan biosynthetic process / HS-GAG biosynthesis / glycosaminoglycan biosynthetic process / lymphocyte migration into lymphoid organs / chondrocyte proliferation /  sulfation / hematopoietic stem cell homeostasis / podocyte differentiation / glandular epithelial cell differentiation / dendritic cell migration / endochondral bone morphogenesis / polysaccharide biosynthetic process / sodium ion homeostasis / endochondral bone growth / acetylglucosaminyltransferase activity / response to light intensity / basement membrane organization / cartilage development involved in endochondral bone morphogenesis / cranial skeletal system development / vocalization behavior / stem cell division / endoderm development / multicellular organismal-level water homeostasis / leukocyte tethering or rolling / vacuole organization / olfactory bulb development / sulfation / hematopoietic stem cell homeostasis / podocyte differentiation / glandular epithelial cell differentiation / dendritic cell migration / endochondral bone morphogenesis / polysaccharide biosynthetic process / sodium ion homeostasis / endochondral bone growth / acetylglucosaminyltransferase activity / response to light intensity / basement membrane organization / cartilage development involved in endochondral bone morphogenesis / cranial skeletal system development / vocalization behavior / stem cell division / endoderm development / multicellular organismal-level water homeostasis / leukocyte tethering or rolling / vacuole organization / olfactory bulb development /  endochondral ossification / fear response / optic nerve development / protein N-linked glycosylation / cellular response to fibroblast growth factor stimulus / neural crest cell differentiation / collagen fibril organization / ossification involved in bone maturation / endochondral ossification / fear response / optic nerve development / protein N-linked glycosylation / cellular response to fibroblast growth factor stimulus / neural crest cell differentiation / collagen fibril organization / ossification involved in bone maturation /  motor behavior / cell adhesion mediated by integrin / epithelial tube branching involved in lung morphogenesis / hair follicle morphogenesis / motor behavior / cell adhesion mediated by integrin / epithelial tube branching involved in lung morphogenesis / hair follicle morphogenesis /  heart contraction / heart contraction /  glycosyltransferase activity / protein glycosylation / glycosyltransferase activity / protein glycosylation /  social behavior / mesoderm development / mesoderm formation / social behavior / mesoderm development / mesoderm formation /  catalytic complex / antigen processing and presentation / canonical Wnt signaling pathway / catalytic complex / antigen processing and presentation / canonical Wnt signaling pathway /  gastrulation / hematopoietic stem cell differentiation / fibroblast growth factor receptor signaling pathway / blood vessel remodeling / gastrulation / hematopoietic stem cell differentiation / fibroblast growth factor receptor signaling pathway / blood vessel remodeling /  cell fate commitment / chondrocyte differentiation / BMP signaling pathway / cell fate commitment / chondrocyte differentiation / BMP signaling pathway /  bone resorption / bone resorption /  ossification / ossification /  synaptic transmission, glutamatergic / synaptic transmission, glutamatergic /  axon guidance / protein catabolic process / multicellular organism growth / axon guidance / protein catabolic process / multicellular organism growth /  wound healing / cellular response to virus / wound healing / cellular response to virus /  regulation of blood pressure / regulation of blood pressure /  vasodilation / vasodilation /  gene expression / protein-containing complex assembly / protein heterodimerization activity gene expression / protein-containing complex assembly / protein heterodimerization activitySimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
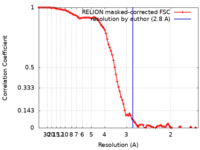
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.8 Å cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Li H | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Chem Biol / Year: 2023 Journal: Nat Chem Biol / Year: 2023Title: Structural basis for heparan sulfate co-polymerase action by the EXT1-2 complex. Authors: Hua Li / Digantkumar Chapla / Robert A Amos / Annapoorani Ramiah / Kelley W Moremen / Huilin Li /  Abstract: Heparan sulfate (HS) proteoglycans are extended (-GlcAβ1,4GlcNAcα1,4-) co-polymers containing decorations of sulfation and epimerization that are linked to cell surface and extracellular matrix ...Heparan sulfate (HS) proteoglycans are extended (-GlcAβ1,4GlcNAcα1,4-) co-polymers containing decorations of sulfation and epimerization that are linked to cell surface and extracellular matrix proteins. In mammals, HS repeat units are extended by an obligate heterocomplex of two exostosin family members, EXT1 and EXT2, where each protein monomer contains distinct GT47 (GT-B fold) and GT64 (GT-A fold) glycosyltransferase domains. In this study, we generated human EXT1-EXT2 (EXT1-2) as a functional heterocomplex and determined its structure in the presence of bound donor and acceptor substrates. Structural data and enzyme activity of catalytic site mutants demonstrate that only two of the four glycosyltransferase domains are major contributors to co-polymer syntheses: the EXT1 GT-B fold β1,4GlcA transferase domain and the EXT2 GT-A fold α1,4GlcNAc transferase domain. The two catalytic sites are over 90 Å apart, indicating that HS is synthesized by a dissociative process that involves a single catalytic site on each monomer. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25037.map.gz emd_25037.map.gz | 56.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25037-v30.xml emd-25037-v30.xml emd-25037.xml emd-25037.xml | 21.9 KB 21.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25037_fsc.xml emd_25037_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_25037.png emd_25037.png | 162.6 KB | ||
| Masks |  emd_25037_msk_1.map emd_25037_msk_1.map | 64 MB |  Mask map Mask map | |
| Others |  emd_25037_additional_1.map.gz emd_25037_additional_1.map.gz emd_25037_half_map_1.map.gz emd_25037_half_map_1.map.gz emd_25037_half_map_2.map.gz emd_25037_half_map_2.map.gz | 6.4 MB 49.7 MB 49.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25037 http://ftp.pdbj.org/pub/emdb/structures/EMD-25037 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25037 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25037 | HTTPS FTP |
-Related structure data
| Related structure data |  7sckMC  7schC  7scjC  7uqxC  7uqyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25037.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25037.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.828 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25037_msk_1.map emd_25037_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_25037_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_25037_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_25037_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human EXT1/2
| Entire | Name: human EXT1/2 |
|---|---|
| Components |
|
-Supramolecule #1: human EXT1/2
| Supramolecule | Name: human EXT1/2 / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Exostosin-1
| Macromolecule | Name: Exostosin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number:  glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 83.404023 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GFRASRSHSR REEHSGRNGL HHPSPDHFWP RFPDALRPFV PWDQLENEDS SVHISPRQKR DANSSIYKGK KCRMESCFDF TLCKKNGFK VYVYPQQKGE KIAESYQNIL AAIEGSRFYT SDPSQACLFV LSLDTLDRDQ LSPQYVHNLR SKVQSLHLWN N GRNHLIFN ...String: GFRASRSHSR REEHSGRNGL HHPSPDHFWP RFPDALRPFV PWDQLENEDS SVHISPRQKR DANSSIYKGK KCRMESCFDF TLCKKNGFK VYVYPQQKGE KIAESYQNIL AAIEGSRFYT SDPSQACLFV LSLDTLDRDQ LSPQYVHNLR SKVQSLHLWN N GRNHLIFN LYSGTWPDYT EDVGFDIGQA MLAKASISTE NFRPNFDVSI PLFSKDHPRT GGERGFLKFN TIPPLRKYML VF KGKRYLT GIGSDTRNAL YHVHNGEDVV LLTTCKHGKD WQKHKDSRCD RDNTEYEKYD YREMLHNATF CLVPRGRRLG SFR FLEALQ AACVPVMLSN GWELPFSEVI NWNQAAVIGD ERLLLQIPST IRSIHQDKIL ALRQQTQFLW EAYFSSVEKI VLTT LEIIQ DRIFKHISRN SLIWNKHPGG LFVLPQYSSY LGDFPYYYAN LGLKPPSKFT AVIHAVTPLV SQSQPVLKLL VAAAK SQYC AQIIVLWNCD KPLPAKHRWP ATAVPVVVIE GESKVMSSRF LPYDNIITDA VLSLDEDTVL STTEVDFAFT VWQSFP ERI VGYPARSHFW DNSKERWGYT SKWTNDYSMV LTGAAIYHKY YHYLYSHYLP ASLKNMVDQL ANCEDILMNF LVSAVTK LP PIKVTQKKQY KETMMGQTSR ASRWADPDHF AQRQSCMNTF ASWFGYMPLI HSQMRLDPVL FKDQVSILRK KYRDIERL |
-Macromolecule #2: Exostosin-2
| Macromolecule | Name: Exostosin-2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO EC number:  glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 77.238031 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GWPHSIESSN DWNVEKRSIR DVPVVRLPAD SPIPERGDLS CRMHTCFDVY RCGFNPKNKI KVYIYALKKY VDDFGVSVSN TISREYNEL LMAISDSDYY TDDINRACLF VPSIDVLNQN TLRIKETAQA MAQLSRWDRG TNHLLFNMLP GGPPDYNTAL D VPRDRALL ...String: GWPHSIESSN DWNVEKRSIR DVPVVRLPAD SPIPERGDLS CRMHTCFDVY RCGFNPKNKI KVYIYALKKY VDDFGVSVSN TISREYNEL LMAISDSDYY TDDINRACLF VPSIDVLNQN TLRIKETAQA MAQLSRWDRG TNHLLFNMLP GGPPDYNTAL D VPRDRALL AGGGFSTWTY RQGYDVSIPV YSPLSAEVDL PEKGPGPRQY FLLSSQVGLH PEYREDLEAL QVKHGESVLV LD KCTNLSE GVLSVRKRCH KHQVFDYPQV LQEATFCVVL RGARLGQAVL SDVLQAGCVP VVIADSYILP FSEVLDWKRA SVV VPEEKM SDVYSILQSI PQRQIEEMQR QARWFWEAYF QSIKAIALAT LQIINDRIYP YAAISYEEWN DPPAVKWGSV SNPL FLPLI PPQSQGFTAI VLTYDRVESL FRVITEVSKV PSLSKLLVVW NNQNKNPPED SLWPKIRVPL KVVRTAENKL SNRFF PYDE IETEAVLAID DDIIMLTSDE LQFGYEVWRE FPDRLVGYPG RLHLWDHEMN KWKYESEWTN EVSMVLTGAA FYHKYF NYL YTYKMPGDIK NWVDAHMNCE DIAMNFLVAN VTGKAVIKVT PRKKFKCPEC TAIDGLSLDQ THMVERSECI NKFASVF GT MPLKVVEHRA DPVLYKDDFP EKLKSFPNIG SL |
-Macromolecule #5: URIDINE-5'-DIPHOSPHATE
| Macromolecule | Name: URIDINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 2 / Formula: UDP |
|---|---|
| Molecular weight | Theoretical: 404.161 Da |
| Chemical component information |  ChemComp-UDP: |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #7: MANGANESE (II) ION
| Macromolecule | Name: MANGANESE (II) ION / type: ligand / ID: 7 / Number of copies: 1 / Formula: MN |
|---|---|
| Molecular weight | Theoretical: 54.938 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7 Component:
| |||||||||
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 299 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 105000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Temperature | Min: 193.0 K / Max: 193.0 K |
| Alignment procedure | Coma free - Residual tilt: 0.05 mrad |
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: GATAN K3 (6k x 4k) / #0 - Number grids imaged: 1 / #0 - Number real images: 5108 / #0 - Average exposure time: 1.5 sec. / #0 - Average electron dose: 68.0 e/Å2 / #1 - Image recording ID: 2 / #1 - Film or detector model: GATAN K3 (6k x 4k) / #1 - Digitization - Dimensions - Width: 5760 pixel / #1 - Digitization - Dimensions - Height: 4092 pixel / #1 - Number grids imaged: 1 / #1 - Number real images: 7609 / #1 - Average exposure time: 1.5 sec. / #1 - Average electron dose: 72.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7sck: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)