+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23609 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PRMT5 bound to covalent PBM-site inhibitor BRD-6988 | |||||||||
 Map data Map data | Map autosharpened by Phenix | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of adenylate cyclase-inhibiting dopamine receptor signaling pathway / peptidyl-arginine N-methylation / oocyte axis specification / type II protein arginine methyltransferase / protein-arginine omega-N symmetric methyltransferase activity / peptidyl-arginine methylation / Golgi ribbon formation / negative regulation of epithelial cell proliferation involved in prostate gland development / histone H4R3 methyltransferase activity / secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development ...positive regulation of adenylate cyclase-inhibiting dopamine receptor signaling pathway / peptidyl-arginine N-methylation / oocyte axis specification / type II protein arginine methyltransferase / protein-arginine omega-N symmetric methyltransferase activity / peptidyl-arginine methylation / Golgi ribbon formation / negative regulation of epithelial cell proliferation involved in prostate gland development / histone H4R3 methyltransferase activity / secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development / epithelial cell proliferation involved in prostate gland development / histone arginine N-methyltransferase activity / methylosome / protein-arginine N-methyltransferase activity / positive regulation of mRNA splicing, via spliceosome / methyl-CpG binding / : /  endothelial cell activation / histone H3 methyltransferase activity / Cul4B-RING E3 ubiquitin ligase complex / endothelial cell activation / histone H3 methyltransferase activity / Cul4B-RING E3 ubiquitin ligase complex /  histone methyltransferase complex / regulation of mitotic nuclear division / positive regulation of oligodendrocyte differentiation / histone methyltransferase complex / regulation of mitotic nuclear division / positive regulation of oligodendrocyte differentiation /  histone methyltransferase activity / histone methyltransferase activity /  E-box binding / negative regulation of cell differentiation / ubiquitin-like ligase-substrate adaptor activity / spliceosomal snRNP assembly / ribonucleoprotein complex binding / regulation of ERK1 and ERK2 cascade / nuclear receptor coactivator activity / regulation of signal transduction by p53 class mediator / E-box binding / negative regulation of cell differentiation / ubiquitin-like ligase-substrate adaptor activity / spliceosomal snRNP assembly / ribonucleoprotein complex binding / regulation of ERK1 and ERK2 cascade / nuclear receptor coactivator activity / regulation of signal transduction by p53 class mediator /  liver regeneration / liver regeneration /  methyltransferase activity / DNA-templated transcription termination / circadian regulation of gene expression / Regulation of TP53 Activity through Methylation / RMTs methylate histone arginines / protein polyubiquitination / transcription corepressor activity / methyltransferase activity / DNA-templated transcription termination / circadian regulation of gene expression / Regulation of TP53 Activity through Methylation / RMTs methylate histone arginines / protein polyubiquitination / transcription corepressor activity /  p53 binding / p53 binding /  snRNP Assembly / ubiquitin-dependent protein catabolic process / snRNP Assembly / ubiquitin-dependent protein catabolic process /  chromatin remodeling / protein heterodimerization activity / positive regulation of cell population proliferation / chromatin remodeling / protein heterodimerization activity / positive regulation of cell population proliferation /  chromatin / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / chromatin / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II /  Golgi apparatus / Golgi apparatus /  nucleoplasm / identical protein binding / nucleoplasm / identical protein binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
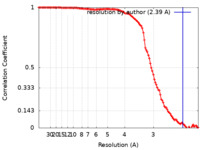
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.39 Å cryo EM / Resolution: 2.39 Å | |||||||||
 Authors Authors | McMillan BJ / McKinney DC / Timm DE | |||||||||
 Citation Citation |  Journal: J Med Chem / Year: 2021 Journal: J Med Chem / Year: 2021Title: Discovery of a First-in-Class Inhibitor of the PRMT5-Substrate Adaptor Interaction. Authors: David C McKinney / Brian J McMillan / Matthew J Ranaghan / Jamie A Moroco / Merissa Brousseau / Zachary Mullin-Bernstein / Meghan O'Keefe / Patrick McCarren / Michael F Mesleh / Kathleen M ...Authors: David C McKinney / Brian J McMillan / Matthew J Ranaghan / Jamie A Moroco / Merissa Brousseau / Zachary Mullin-Bernstein / Meghan O'Keefe / Patrick McCarren / Michael F Mesleh / Kathleen M Mulvaney / Foxy Robinson / Ritu Singh / Besnik Bajrami / Florence F Wagner / Robert Hilgraf / Martin J Drysdale / Arthur J Campbell / Adam Skepner / David E Timm / Dale Porter / Virendar K Kaushik / William R Sellers / Alessandra Ianari /  Abstract: PRMT5 and its substrate adaptor proteins (SAPs), pICln and Riok1, are synthetic lethal dependencies in MTAP-deleted cancer cells. SAPs share a conserved PRMT5 binding motif (PBM) which mediates ...PRMT5 and its substrate adaptor proteins (SAPs), pICln and Riok1, are synthetic lethal dependencies in MTAP-deleted cancer cells. SAPs share a conserved PRMT5 binding motif (PBM) which mediates binding to a surface of PRMT5 distal to the catalytic site. This interaction is required for methylation of several PRMT5 substrates, including histone and spliceosome complexes. We screened for small molecule inhibitors of the PRMT5-PBM interaction and validated a compound series which binds to the PRMT5-PBM interface and directly inhibits binding of SAPs. Mode of action studies revealed the formation of a covalent bond between a halogenated pyridazinone group and cysteine 278 of PRMT5. Optimization of the starting hit produced a lead compound, BRD0639, which engages the target in cells, disrupts PRMT5-RIOK1 complexes, and reduces substrate methylation. BRD0639 is a first-in-class PBM-competitive inhibitor that can support studies of PBM-dependent PRMT5 activities and the development of novel PRMT5 inhibitors that selectively target these functions. #1:  Journal: bioRxiv / Year: 2020 Journal: bioRxiv / Year: 2020Title: Discovery of a first-in-class inhibitor of the PRMT5-substrate adaptor interaction Authors: Mulvaney KM / McMillan BJ / Sellers WR | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23609.map.gz emd_23609.map.gz | 116.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23609-v30.xml emd-23609-v30.xml emd-23609.xml emd-23609.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23609_fsc.xml emd_23609_fsc.xml | 14.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_23609.png emd_23609.png | 124 KB | ||
| Masks |  emd_23609_msk_1.map emd_23609_msk_1.map | 7.5 MB |  Mask map Mask map | |
| Others |  emd_23609_half_map_1.map.gz emd_23609_half_map_1.map.gz emd_23609_half_map_2.map.gz emd_23609_half_map_2.map.gz | 11.4 MB 11.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23609 http://ftp.pdbj.org/pub/emdb/structures/EMD-23609 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23609 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23609 | HTTPS FTP |
-Related structure data
| Related structure data |  7m05MC  6v0pC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23609.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23609.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map autosharpened by Phenix | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_23609_msk_1.map emd_23609_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
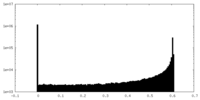
| Density Histograms |
-Half map: cisTEM half-map 2
| File | emd_23609_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cisTEM half-map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
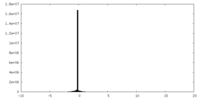
| Density Histograms |
-Half map: cisTEM half-map 1
| File | emd_23609_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cisTEM half-map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
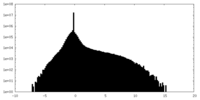
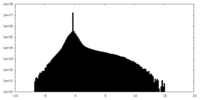
| Density Histograms |
- Sample components
Sample components
-Entire : Hetero-octamer complex of PRMT5 and WDR77
| Entire | Name: Hetero-octamer complex of PRMT5 and WDR77 |
|---|---|
| Components |
|
-Supramolecule #1: Hetero-octamer complex of PRMT5 and WDR77
| Supramolecule | Name: Hetero-octamer complex of PRMT5 and WDR77 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Molecular weight | Theoretical: 437 KDa |
-Macromolecule #1: Protein arginine N-methyltransferase 5
| Macromolecule | Name: Protein arginine N-methyltransferase 5 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: type II protein arginine methyltransferase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 72.766664 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MAAMAVGGAG GSRVSSGRDL NCVPEIADTL GAVAKQGFDF LCMPVFHPRF KREFIQEPAK NRPGPQTRSD LLLSGRDWNT LIVGKLSPW IRPDSKVEKI RRNSEAAMLQ ELNFGAYLGL PAFLLPLNQE DNTNLARVLT NHIHTGHHSS MFWMRVPLVA P EDLRDDII ...String: MAAMAVGGAG GSRVSSGRDL NCVPEIADTL GAVAKQGFDF LCMPVFHPRF KREFIQEPAK NRPGPQTRSD LLLSGRDWNT LIVGKLSPW IRPDSKVEKI RRNSEAAMLQ ELNFGAYLGL PAFLLPLNQE DNTNLARVLT NHIHTGHHSS MFWMRVPLVA P EDLRDDII ENAPTTHTEE YSGEEKTWMW WHNFRTLCDY SKRIAVALEI GADLPSNHVI DRWLGEPIKA AILPTSIFLT NK KGFPVLS KMHQRLIFRL LKLEVQFIIT GTNHHSEKEF CSYLQYLEYL SQNRPPPNAY ELFAKGYEDY LQSPLQPLMD NLE SQTYEV FEKDPIKYSQ YQQAIYKCLL DRVPEEEKDT NVQVLMVLGA GRGPLVNASL RAAKQADRRI KLYAVEKNPN AVVT LENWQ FEEWGSQVTV VSSDMREWVA PEKADIIVSE LLGSFADNEL SPECLDGAQH FLKDDGVSIP GEYTSFLAPI SSSKL YNEV RACREKDRDP EAQFEMPYVV RLHNFHQLSA PQPCFTFSHP NRDPMIDNNR YCTLEFPVEV NTVLHGFAGY FETVLY QDI TLSIRPETHS PGMFSWFPIL FPIKQPITVR EGQTICVRFW RCSNSKKVWY EWAVTAPVCS AIHNPTGRSY TIGL |
-Macromolecule #2: Methylosome protein 50
| Macromolecule | Name: Methylosome protein 50 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 36.723164 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: PRKETPPPLV PPAAREWNLP PNAPACMERQ LEAARYRSDG ALLLGASSLS GRCWAGSLWL FKDPCAAPNE GFCSAGVQTE AGVADLTWV GERGILVASD SGAVELWELD ENETLIVSKF CKYEHDDIVS TVSVLSSGTQ AVSGSKDICI KVWDLAQQVV L SSYRAHAA ...String: PRKETPPPLV PPAAREWNLP PNAPACMERQ LEAARYRSDG ALLLGASSLS GRCWAGSLWL FKDPCAAPNE GFCSAGVQTE AGVADLTWV GERGILVASD SGAVELWELD ENETLIVSKF CKYEHDDIVS TVSVLSSGTQ AVSGSKDICI KVWDLAQQVV L SSYRAHAA QVTCVAASPH KDSVFLSCSE DNRILLWDTR CPKPASQIGC SAPGYLPTSL AWHPQQSEVF VFGDENGTVS LV DTKSTSC VLSSAVHSQC VTGLVFSPHS VPFLASLSED CSLAVLDSSL SELFRSQAHR DFVRDATWSP LNHSLLTTVG WDH QVVHHV VPTEPLPAPG PASVTE |
-Macromolecule #3: 2-(5-chloro-6-oxopyridazin-1(6H)-yl)-N-(4-methyl-3-{[2-(pyridin-2...
| Macromolecule | Name: 2-(5-chloro-6-oxopyridazin-1(6H)-yl)-N-(4-methyl-3-{[2-(pyridin-2-yl)ethyl]sulfamoyl}phenyl)acetamide type: ligand / ID: 3 / Number of copies: 4 / Formula: YJG |
|---|---|
| Molecular weight | Theoretical: 461.922 Da |
| Chemical component information | 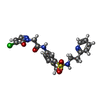 ChemComp-YJG: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.42 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||
| Grid | Model: UltrAuFoil / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 46296 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 1.4000000000000001 µm / Nominal magnification: 130000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 1.4000000000000001 µm / Nominal magnification: 130000 |
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 30 eV |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Sampling interval: 5.0 µm / Digitization - Frames/image: 4-40 / Number grids imaged: 1 / Number real images: 2169 / Average exposure time: 8.0 sec. / Average electron dose: 62.4 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z
Z Y
Y X
X