+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
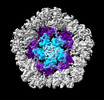
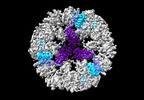
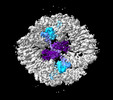
| Title | Structure of the PNMA2 capsid | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Endogenous retrovirus. PNMA2 / PNMA / Paraneoplastic syndrome / Paraneoplastic antigen Ma2 / VLP. /  VIRUS LIKE PARTICLE VIRUS LIKE PARTICLE | ||||||||||||
| Function / homology |  : / : /  : / PNMA N-terminal RRM-like domain / Paraneoplastic antigen Ma / PNMA / : / PNMA N-terminal RRM-like domain / Paraneoplastic antigen Ma / PNMA /  nucleolus / Paraneoplastic antigen Ma2 homolog nucleolus / Paraneoplastic antigen Ma2 homolog Function and homology information Function and homology information | ||||||||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.2 Å cryo EM / Resolution: 3.2 Å | ||||||||||||
 Authors Authors | Erlendsson S / Xu J / Shepherd JD / Briggs JAG | ||||||||||||
| Funding support |  United States, United States,  United Kingdom, United Kingdom,  Denmark, 3 items Denmark, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: PNMA2 forms immunogenic non-enveloped virus-like capsids associated with paraneoplastic neurological syndrome. Authors: Junjie Xu / Simon Erlendsson / Manvendra Singh / G Aaron Holling / Matthew Regier / Iosune Ibiricu / Jenifer Einstein / Michael P Hantak / Gregory S Day / Amanda L Piquet / Tammy L Smith / ...Authors: Junjie Xu / Simon Erlendsson / Manvendra Singh / G Aaron Holling / Matthew Regier / Iosune Ibiricu / Jenifer Einstein / Michael P Hantak / Gregory S Day / Amanda L Piquet / Tammy L Smith / Stacey L Clardy / Alexandra M Whiteley / Cédric Feschotte / John A G Briggs / Jason D Shepherd /    Abstract: The paraneoplastic Ma antigen (PNMA) proteins are associated with cancer-induced paraneoplastic syndromes that present with an autoimmune response and neurological symptoms. Why PNMA proteins are ...The paraneoplastic Ma antigen (PNMA) proteins are associated with cancer-induced paraneoplastic syndromes that present with an autoimmune response and neurological symptoms. Why PNMA proteins are associated with this severe autoimmune disease is unclear. PNMA genes are predominantly expressed in the central nervous system and are ectopically expressed in some tumors. We show that PNMA2, which has been co-opted from a Ty3 retrotransposon, encodes a protein that is released from cells as non-enveloped virus-like capsids. Recombinant PNMA2 capsids injected into mice induce autoantibodies that preferentially bind external "spike" PNMA2 capsid epitopes, whereas a capsid-assembly-defective PNMA2 protein is not immunogenic. PNMA2 autoantibodies in cerebrospinal fluid of patients with anti-Ma2 paraneoplastic disease show similar preferential binding to spike capsid epitopes. PNMA2 capsid-injected mice develop learning and memory deficits. These observations suggest that PNMA2 capsids act as an extracellular antigen, capable of generating an autoimmune response that results in neurological deficits. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19024.map.gz emd_19024.map.gz | 45.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19024-v30.xml emd-19024-v30.xml emd-19024.xml emd-19024.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_19024.png emd_19024.png | 64 KB | ||
| Filedesc metadata |  emd-19024.cif.gz emd-19024.cif.gz | 6.3 KB | ||
| Others |  emd_19024_additional_1.map.gz emd_19024_additional_1.map.gz emd_19024_half_map_1.map.gz emd_19024_half_map_1.map.gz emd_19024_half_map_2.map.gz emd_19024_half_map_2.map.gz | 405.4 MB 407.3 MB 407.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19024 http://ftp.pdbj.org/pub/emdb/structures/EMD-19024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19024 | HTTPS FTP |
-Related structure data
| Related structure data |  8rb3MC  8rb4C  8rb5C  8rb7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19024.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19024.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_19024_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
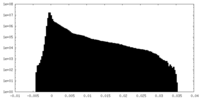
| Density Histograms |
-Half map: #2
| File | emd_19024_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_19024_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of the PNMA2 capsid
| Entire | Name: Structure of the PNMA2 capsid |
|---|---|
| Components |
|
-Supramolecule #1: Structure of the PNMA2 capsid
| Supramolecule | Name: Structure of the PNMA2 capsid / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: T=1 icosahedral mPNMA2 capsid consisting of 60 PNMA2 molecules |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
-Macromolecule #1: Paraneoplastic antigen Ma2 homolog
| Macromolecule | Name: Paraneoplastic antigen Ma2 homolog / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Molecular weight | Theoretical: 20.667789 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: LLPVKYCKMR IFSGSTAAAP EEEPFEVWLE QATEIAKEWP IPEAEKKRWV AESLRGPALD LMHIVQADNP SISVGECLEA FKQVFGSTE SRRTSQVKYL RTYQQEGEKI SAYVLRLETL LRRAVEKRAI PRNIADQVRL EQVMAGANLG NVLWCRLQEL K DQGPLPTF LQLMKVIREE EE UniProtKB: Paraneoplastic antigen Ma2 homolog |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 35 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 96000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number real images: 3005 / Average exposure time: 40.0 sec. / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 47652 |
|---|---|
| Startup model | Type of model: NONE |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final 3D classification | Number classes: 4 / Software - Name: RELION (ver. 4) Details: Proceeded with 90 percent of capsids af classification |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 4) |
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: I (icosahedral ) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 4) / Number images used: 35420 ) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 4) / Number images used: 35420 |
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Space: REAL / Overall B value: 147 |
| Output model |  PDB-8rb3: |
 Movie
Movie Controller
Controller



 Z
Z Y
Y X
X