+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human apo ALDH1A1 determined by cryoEM at 100 keV | |||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | Oxidoreductase Aldehyde dehydrogenase /  OXIDOREDUCTASE OXIDOREDUCTASE | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationfructosamine catabolic process / 3-deoxyglucosone dehydrogenase activity /  benzaldehyde dehydrogenase (NAD+) / benzaldehyde dehydrogenase (NAD+) /  benzaldehyde dehydrogenase (NAD+) activity / maintenance of lens transparency / gamma-aminobutyric acid biosynthetic process / benzaldehyde dehydrogenase (NAD+) activity / maintenance of lens transparency / gamma-aminobutyric acid biosynthetic process /  aminobutyraldehyde dehydrogenase activity / aminobutyraldehyde dehydrogenase activity /  retinal dehydrogenase / retinal dehydrogenase /  aminobutyraldehyde dehydrogenase / Fructose catabolism ...fructosamine catabolic process / 3-deoxyglucosone dehydrogenase activity / aminobutyraldehyde dehydrogenase / Fructose catabolism ...fructosamine catabolic process / 3-deoxyglucosone dehydrogenase activity /  benzaldehyde dehydrogenase (NAD+) / benzaldehyde dehydrogenase (NAD+) /  benzaldehyde dehydrogenase (NAD+) activity / maintenance of lens transparency / gamma-aminobutyric acid biosynthetic process / benzaldehyde dehydrogenase (NAD+) activity / maintenance of lens transparency / gamma-aminobutyric acid biosynthetic process /  aminobutyraldehyde dehydrogenase activity / aminobutyraldehyde dehydrogenase activity /  retinal dehydrogenase / retinal dehydrogenase /  aminobutyraldehyde dehydrogenase / Fructose catabolism / cellular aldehyde metabolic process / Ethanol oxidation / RA biosynthesis pathway / glyceraldehyde-3-phosphate dehydrogenase (NAD+) (non-phosphorylating) activity / aminobutyraldehyde dehydrogenase / Fructose catabolism / cellular aldehyde metabolic process / Ethanol oxidation / RA biosynthesis pathway / glyceraldehyde-3-phosphate dehydrogenase (NAD+) (non-phosphorylating) activity /  aldehyde dehydrogenase (NAD+) / cellular detoxification of aldehyde / aldehyde dehydrogenase (NAD+) / cellular detoxification of aldehyde /  androgen binding / androgen binding /  aldehyde dehydrogenase (NAD+) activity / negative regulation of cold-induced thermogenesis / aldehyde dehydrogenase (NAD+) activity / negative regulation of cold-induced thermogenesis /  retinal dehydrogenase activity / retinol metabolic process / retinoid metabolic process / retinal dehydrogenase activity / retinol metabolic process / retinoid metabolic process /  GTPase activator activity / NAD binding / GTPase activator activity / NAD binding /  axon / axon /  synapse / extracellular exosome / synapse / extracellular exosome /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||||||||||||||
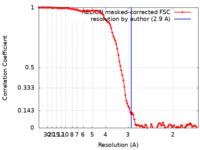
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.9 Å cryo EM / Resolution: 2.9 Å | |||||||||||||||||||||
 Authors Authors | McMullan G / Naydenova K / Mihaylov D / Peet MJ / Wilson H / Yamashita K / Dickerson JL / Chen S / Cannone G / Lee Y ...McMullan G / Naydenova K / Mihaylov D / Peet MJ / Wilson H / Yamashita K / Dickerson JL / Chen S / Cannone G / Lee Y / Hutchings KA / Gittins O / Sobhy M / Wells T / El-Gomati MM / Dalby J / Meffert M / Schulze-Briese C / Henderson R / Russo CJ | |||||||||||||||||||||
| Funding support |  United Kingdom, 6 items United Kingdom, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structure determination by cryoEM at 100 keV. Authors: Greg McMullan / Katerina Naydenova / Daniel Mihaylov / Keitaro Yamashita / Mathew J Peet / Hugh Wilson / Joshua L Dickerson / Shaoxia Chen / Giuseppe Cannone / Yang Lee / Katherine A ...Authors: Greg McMullan / Katerina Naydenova / Daniel Mihaylov / Keitaro Yamashita / Mathew J Peet / Hugh Wilson / Joshua L Dickerson / Shaoxia Chen / Giuseppe Cannone / Yang Lee / Katherine A Hutchings / Olivia Gittins / Mohamed A Sobhy / Torquil Wells / Mohamed M El-Gomati / Jason Dalby / Matthias Meffert / Clemens Schulze-Briese / Richard Henderson / Christopher J Russo /    Abstract: Electron cryomicroscopy can, in principle, determine the structures of most biological molecules but is currently limited by access, specimen preparation difficulties, and cost. We describe a purpose- ...Electron cryomicroscopy can, in principle, determine the structures of most biological molecules but is currently limited by access, specimen preparation difficulties, and cost. We describe a purpose-built instrument operating at 100 keV-including advances in electron optics, detection, and processing-that makes structure determination fast and simple at a fraction of current costs. The instrument attains its theoretical performance limits, allowing atomic resolution imaging of gold test specimens and biological molecular structure determination in hours. We demonstrate its capabilities by determining the structures of eleven different specimens, ranging in size from 140 kDa to 2 MDa, using a fraction of the data normally required. CryoEM with a microscope designed specifically for high-efficiency, on-the-spot imaging of biological molecules will expand structural biology to a wide range of previously intractable problems. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17966.map.gz emd_17966.map.gz | 20.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17966-v30.xml emd-17966-v30.xml emd-17966.xml emd-17966.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17966_fsc.xml emd_17966_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_17966.png emd_17966.png | 100.7 KB | ||
| Masks |  emd_17966_msk_1.map emd_17966_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17966.cif.gz emd-17966.cif.gz | 6.3 KB | ||
| Others |  emd_17966_half_map_1.map.gz emd_17966_half_map_1.map.gz emd_17966_half_map_2.map.gz emd_17966_half_map_2.map.gz | 97.2 MB 96.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17966 http://ftp.pdbj.org/pub/emdb/structures/EMD-17966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17966 | HTTPS FTP |
-Related structure data
| Related structure data |  8pvhMC  8pv9C  8pvaC  8pvbC  8pvcC  8pvdC  8pveC  8pvfC  8pvgC  8pviC  8pvjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17966.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17966.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.8415 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17966_msk_1.map emd_17966_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17966_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17966_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human apo ALDH1A1
| Entire | Name: Human apo ALDH1A1 |
|---|---|
| Components |
|
-Supramolecule #1: Human apo ALDH1A1
| Supramolecule | Name: Human apo ALDH1A1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Aldehyde dehydrogenase 1A1
| Macromolecule | Name: Aldehyde dehydrogenase 1A1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number:  aminobutyraldehyde dehydrogenase aminobutyraldehyde dehydrogenase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 57.99298 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MSYYHHHHHH LESTSLYKKA GSAAAPFTSS SGTPDLPVLL TDLKIQYTKI FINNEWHDSV SGKKFPVFNP ATEEELCQVE EGDKEDVDK AVKAARQAFQ IGSPWRTMDA SERGRLLYKL ADLIERDRLL LATMESMNGG KLYSNAYLND LAGCIKTLRY C AGWADKIQ ...String: MSYYHHHHHH LESTSLYKKA GSAAAPFTSS SGTPDLPVLL TDLKIQYTKI FINNEWHDSV SGKKFPVFNP ATEEELCQVE EGDKEDVDK AVKAARQAFQ IGSPWRTMDA SERGRLLYKL ADLIERDRLL LATMESMNGG KLYSNAYLND LAGCIKTLRY C AGWADKIQ GRTIPIDGNF FTYTRHEPIG VCGQIIPWNF PLVMLIWKIG PALSCGNTVV VKPAEQTPLT ALHVASLIKE AG FPPGVVN IVPGYGPTAG AAISSHMDID KVAFTGSTEV GKLIKEAAGK SNLKRVTLEL GGKSPCIVLA DADLDNAVEF AHH GVFYHQ GQCCIAASRI FVEESIYDEF VRRSVERAKK YILGNPLTPG VTQGPQIDKE QYDKILDLIE SGKKEGAKLE CGGG PWGNK GYFVQPTVFS NVTDEMRIAK EEIFGPVQQI MKFKSLDDVI KRANNTFYGL SAGVFTKDID KAITISSALQ AGTVW VNCY GVVSAQCPFG GFKMSGNGRE LGEYGFHEYT EVKTVTVKIS QKNS UniProtKB: Aldehyde dehydrogenase 1A1 |
-Macromolecule #2: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 2 / Number of copies: 1 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: UltrAuFoil R0./1 / Material: GOLD |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 1400/HR + YPS FEG |
|---|---|
| Electron beam | Acceleration voltage: 100 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: DECTRIS SINGLA (1k x 1k) / Average electron dose: 41.0 e/Å2 |
 Movie
Movie Controller
Controller


















 Z
Z Y
Y X
X