[English] 日本語
 Yorodumi
Yorodumi- EMDB-15697: Cryo-EM structure of shCas12k-sgRNA-dsDNA ternary complex (type V... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of shCas12k-sgRNA-dsDNA ternary complex (type V-K CRISPR-associated transposon) | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords |  CRISPR / CRISPR /  complex / transposition / complex / transposition /  DNA BINDING PROTEIN DNA BINDING PROTEIN | |||||||||||||||
| Function / homology | Cas12k Function and homology information Function and homology information | |||||||||||||||
| Biological species |  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) | |||||||||||||||
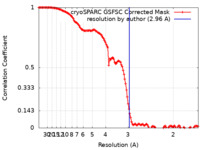
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.96 Å cryo EM / Resolution: 2.96 Å | |||||||||||||||
 Authors Authors | Tenjo-Castano F / Sofos N / Stella S / Fuglsang A / Pape T / Mesa P / Stutzke LS / Temperini P / Montoya G | |||||||||||||||
| Funding support |  Denmark, 4 items Denmark, 4 items
| |||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Conformational Landscape of the Type V-K CRISPR-associated Transposon Integration Assembly Authors: Tenjo-Castano F / Sofos N / Stella S / Fuglsang A / Pape T / Mesa P / Stutzke LS / Temperini P / Montoya G | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15697.map.gz emd_15697.map.gz | 108.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15697-v30.xml emd-15697-v30.xml emd-15697.xml emd-15697.xml | 25.6 KB 25.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15697_fsc.xml emd_15697_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_15697.png emd_15697.png | 53.7 KB | ||
| Masks |  emd_15697_msk_1.map emd_15697_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15697.cif.gz emd-15697.cif.gz | 7 KB | ||
| Others |  emd_15697_additional_1.map.gz emd_15697_additional_1.map.gz emd_15697_half_map_1.map.gz emd_15697_half_map_1.map.gz emd_15697_half_map_2.map.gz emd_15697_half_map_2.map.gz | 195.4 MB 200.7 MB 200.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15697 http://ftp.pdbj.org/pub/emdb/structures/EMD-15697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15697 | HTTPS FTP |
-Related structure data
| Related structure data |  8axaMC  19075  19282  19283  19284  19286 M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15697.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15697.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15697_msk_1.map emd_15697_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
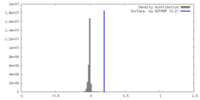
| Density Histograms |
-Additional map: DeepEMhancer map used for model building
| File | emd_15697_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DeepEMhancer map used for model building | ||||||||||||
| Projections & Slices |
| ||||||||||||
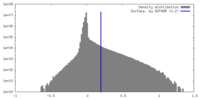
| Density Histograms |
-Half map: #2
| File | emd_15697_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
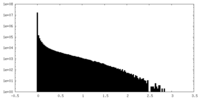
| Density Histograms |
-Half map: #1
| File | emd_15697_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
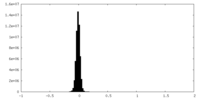
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of ShCas12k bound to sgRNA and dsDNA.
| Entire | Name: Ternary complex of ShCas12k bound to sgRNA and dsDNA. |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of ShCas12k bound to sgRNA and dsDNA.
| Supramolecule | Name: Ternary complex of ShCas12k bound to sgRNA and dsDNA. / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 193 KDa |
-Macromolecule #1: Cas12k
| Macromolecule | Name: Cas12k / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 74.738258 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MSQITIQARL ISFESNRQQL WKLMADLNTP LINELLCQLG QHPDFEKWQQ KGKLPSTVVS QLCQPLKTDP RFAGQPSRLY MSAIHIVDY IYKSWLAIQK RLQQQLDGKT RWLEMLNSDA ELVELSGDTL EAIRVKAAEI LAIAMPASES DSASPKGKKG K KEKKPSSS ...String: MSQITIQARL ISFESNRQQL WKLMADLNTP LINELLCQLG QHPDFEKWQQ KGKLPSTVVS QLCQPLKTDP RFAGQPSRLY MSAIHIVDY IYKSWLAIQK RLQQQLDGKT RWLEMLNSDA ELVELSGDTL EAIRVKAAEI LAIAMPASES DSASPKGKKG K KEKKPSSS SPKRSLSKTL FDAYQETEDI KSRSAISYLL KNGCKLTDKE EDSEKFAKRR RQVEIQIQRL TEKLISRMPK GR DLTNAKW LETLLTATTT VAEDNAQAKR WQDILLTRSS SLPFPLVFET NEDMVWSKNQ KGRLCVHFNG LSDLIFEVYC GNR QLHWFQ RFLEDQQTKR KSKNQHSSGL FTLRNGHLVW LEGEGKGEPW NLHHLTLYCC VDNRLWTEEG TEIVRQEKAD EITK FITNM KKKSDLSDTQ QALIQRKQST LTRINNSFER PSQPLYQGQS HILVGVSLGL EKPATVAVVD AIANKVLAYR SIKQL LGDN YELLNRQRRQ QQYLSHERHK AQKNFSPNQF GASELGQHID RLLAKAIVAL ARTYKAGSIV LPKLGDMREV VQSEIQ AIA EQKFPGYIEG QQKYAKQYRV NVHRWSYGRL IQSIQSKAAQ TGIVIEEGKQ PIRGSPHDKA KELALSAYNL RLTRRSG SE FELENLYFQ UniProtKB: Cas12k |
-Macromolecule #2: sgRNA
| Macromolecule | Name: sgRNA / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 84.006516 KDa |
| Sequence | String: GGAUAUUAAU AGCGCCGCAA UUCAUGCUGC UUGCAGCCUC UGAAUUUUGU UAAAUGAGGG UUAGUUUGAC UGUAUAAAUA CAGUCUUGC UUUCUGACCC UGGUAGCUGC UCACCCUGAU GCUGCUGUCA AUAGACAGGA UAGGUGCGCU CCCAGCAAUA A GGGCGCGG ...String: GGAUAUUAAU AGCGCCGCAA UUCAUGCUGC UUGCAGCCUC UGAAUUUUGU UAAAUGAGGG UUAGUUUGAC UGUAUAAAUA CAGUCUUGC UUUCUGACCC UGGUAGCUGC UCACCCUGAU GCUGCUGUCA AUAGACAGGA UAGGUGCGCU CCCAGCAAUA A GGGCGCGG AUGUACUGCU GUAGUGGCUA CUGAAUCACC CCCGAUCAAG GGGGAACCCU CCAAAAGGUG GGUUGAAAGG AG AAGUCAU UUAAUAAGGC CACU |
-Macromolecule #3: DNA target strand
| Macromolecule | Name: DNA target strand / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 16.858855 KDa |
| Sequence | String: (DC)(DT)(DA)(DG)(DA)(DA)(DA)(DG)(DA)(DC) (DT)(DT)(DT)(DT)(DA)(DA)(DC)(DA)(DG)(DT) (DG)(DG)(DC)(DC)(DT)(DT)(DA)(DT)(DT) (DA)(DA)(DA)(DT)(DG)(DA)(DC)(DT)(DT)(DC) (DT) (DC)(DA)(DA)(DC)(DC)(DA) ...String: (DC)(DT)(DA)(DG)(DA)(DA)(DA)(DG)(DA)(DC) (DT)(DT)(DT)(DT)(DA)(DA)(DC)(DA)(DG)(DT) (DG)(DG)(DC)(DC)(DT)(DT)(DA)(DT)(DT) (DA)(DA)(DA)(DT)(DG)(DA)(DC)(DT)(DT)(DC) (DT) (DC)(DA)(DA)(DC)(DC)(DA)(DT)(DC) (DT)(DT)(DG)(DC)(DT)(DG)(DA) |
-Macromolecule #4: DNA non-target strand
| Macromolecule | Name: DNA non-target strand / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Molecular weight | Theoretical: 17.027963 KDa |
| Sequence | String: (DT)(DC)(DA)(DG)(DC)(DA)(DA)(DG)(DA)(DT) (DG)(DG)(DT)(DT)(DG)(DA)(DG)(DA)(DA)(DG) (DT)(DC)(DA)(DT)(DT)(DT)(DA)(DA)(DT) (DA)(DA)(DG)(DG)(DC)(DC)(DA)(DC)(DT)(DG) (DT) (DT)(DA)(DA)(DA)(DA)(DG) ...String: (DT)(DC)(DA)(DG)(DC)(DA)(DA)(DG)(DA)(DT) (DG)(DG)(DT)(DT)(DG)(DA)(DG)(DA)(DA)(DG) (DT)(DC)(DA)(DT)(DT)(DT)(DA)(DA)(DT) (DA)(DA)(DG)(DG)(DC)(DC)(DA)(DC)(DT)(DG) (DT) (DT)(DA)(DA)(DA)(DA)(DG)(DT)(DC) (DT)(DT)(DT)(DC)(DT)(DA)(DG) |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 3 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 9.6 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 3 s blotting, 4 degrees celcius. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 96000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 2231 / Average exposure time: 45.0 sec. / Average electron dose: 45.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8axa: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)