+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the V2 receptor Cter-arrestin2-ScFv30 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information angiotensin receptor binding / Activation of SMO / negative regulation of interleukin-8 production / arrestin family protein binding / G protein-coupled receptor internalization / angiotensin receptor binding / Activation of SMO / negative regulation of interleukin-8 production / arrestin family protein binding / G protein-coupled receptor internalization /  enzyme inhibitor activity / Lysosome Vesicle Biogenesis / positive regulation of Rho protein signal transduction / Golgi Associated Vesicle Biogenesis / negative regulation of NF-kappaB transcription factor activity ... enzyme inhibitor activity / Lysosome Vesicle Biogenesis / positive regulation of Rho protein signal transduction / Golgi Associated Vesicle Biogenesis / negative regulation of NF-kappaB transcription factor activity ... angiotensin receptor binding / Activation of SMO / negative regulation of interleukin-8 production / arrestin family protein binding / G protein-coupled receptor internalization / angiotensin receptor binding / Activation of SMO / negative regulation of interleukin-8 production / arrestin family protein binding / G protein-coupled receptor internalization /  enzyme inhibitor activity / Lysosome Vesicle Biogenesis / positive regulation of Rho protein signal transduction / Golgi Associated Vesicle Biogenesis / negative regulation of NF-kappaB transcription factor activity / enzyme inhibitor activity / Lysosome Vesicle Biogenesis / positive regulation of Rho protein signal transduction / Golgi Associated Vesicle Biogenesis / negative regulation of NF-kappaB transcription factor activity /  stress fiber assembly / negative regulation of Notch signaling pathway / stress fiber assembly / negative regulation of Notch signaling pathway /  pseudopodium / negative regulation of interleukin-6 production / positive regulation of receptor internalization / pseudopodium / negative regulation of interleukin-6 production / positive regulation of receptor internalization /  clathrin-coated pit / negative regulation of protein ubiquitination / clathrin-coated pit / negative regulation of protein ubiquitination /  insulin-like growth factor receptor binding / insulin-like growth factor receptor binding /  visual perception / Activated NOTCH1 Transmits Signal to the Nucleus / visual perception / Activated NOTCH1 Transmits Signal to the Nucleus /  GTPase activator activity / G protein-coupled receptor binding / Signaling by high-kinase activity BRAF mutants / MAP2K and MAPK activation / cytoplasmic vesicle membrane / Signaling by RAF1 mutants / Signaling by moderate kinase activity BRAF mutants / Paradoxical activation of RAF signaling by kinase inactive BRAF / Signaling downstream of RAS mutants / Signaling by BRAF and RAF1 fusions / Thrombin signalling through proteinase activated receptors (PARs) / GTPase activator activity / G protein-coupled receptor binding / Signaling by high-kinase activity BRAF mutants / MAP2K and MAPK activation / cytoplasmic vesicle membrane / Signaling by RAF1 mutants / Signaling by moderate kinase activity BRAF mutants / Paradoxical activation of RAF signaling by kinase inactive BRAF / Signaling downstream of RAS mutants / Signaling by BRAF and RAF1 fusions / Thrombin signalling through proteinase activated receptors (PARs) /  protein transport / Cargo recognition for clathrin-mediated endocytosis / protein transport / Cargo recognition for clathrin-mediated endocytosis /  Clathrin-mediated endocytosis / ubiquitin-dependent protein catabolic process / cytoplasmic vesicle / G alpha (s) signalling events / proteasome-mediated ubiquitin-dependent protein catabolic process / Clathrin-mediated endocytosis / ubiquitin-dependent protein catabolic process / cytoplasmic vesicle / G alpha (s) signalling events / proteasome-mediated ubiquitin-dependent protein catabolic process /  transcription coactivator activity / positive regulation of ERK1 and ERK2 cascade / protein ubiquitination / transcription coactivator activity / positive regulation of ERK1 and ERK2 cascade / protein ubiquitination /  nuclear body / Ub-specific processing proteases / positive regulation of protein phosphorylation / lysosomal membrane / nuclear body / Ub-specific processing proteases / positive regulation of protein phosphorylation / lysosomal membrane /  Golgi membrane / Golgi membrane /  ubiquitin protein ligase binding / ubiquitin protein ligase binding /  chromatin / regulation of transcription by RNA polymerase II / chromatin / regulation of transcription by RNA polymerase II /  signal transduction / positive regulation of transcription by RNA polymerase II / signal transduction / positive regulation of transcription by RNA polymerase II /  nucleoplasm / nucleoplasm /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 4.23 Å cryo EM / Resolution: 4.23 Å | |||||||||
 Authors Authors | Bous J / Fouillen A / Trapani S / Granier S / Mouillac B / Bron P | |||||||||
| Funding support |  France, 2 items France, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Structure of the vasopressin hormone-V2 receptor-β-arrestin1 ternary complex. Authors: Julien Bous / Aurélien Fouillen / Hélène Orcel / Stefano Trapani / Xiaojing Cong / Simon Fontanel / Julie Saint-Paul / Joséphine Lai-Kee-Him / Serge Urbach / Nathalie Sibille / Rémy ...Authors: Julien Bous / Aurélien Fouillen / Hélène Orcel / Stefano Trapani / Xiaojing Cong / Simon Fontanel / Julie Saint-Paul / Joséphine Lai-Kee-Him / Serge Urbach / Nathalie Sibille / Rémy Sounier / Sébastien Granier / Bernard Mouillac / Patrick Bron /  Abstract: Arrestins interact with G protein-coupled receptors (GPCRs) to stop G protein activation and to initiate key signaling pathways. Recent structural studies shed light on the molecular mechanisms ...Arrestins interact with G protein-coupled receptors (GPCRs) to stop G protein activation and to initiate key signaling pathways. Recent structural studies shed light on the molecular mechanisms involved in GPCR-arrestin coupling, but whether this process is conserved among GPCRs is poorly understood. Here, we report the cryo-electron microscopy active structure of the wild-type arginine-vasopressin V2 receptor (V2R) in complex with β-arrestin1. It reveals an atypical position of β-arrestin1 compared to previously described GPCR-arrestin assemblies, associated with an original V2R/β-arrestin1 interface involving all receptor intracellular loops. Phosphorylated sites of the V2R carboxyl terminus are clearly identified and interact extensively with the β-arrestin1 N-lobe, in agreement with structural data obtained with chimeric or synthetic systems. Overall, these findings highlight a notable structural variability among GPCR-arrestin signaling complexes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14223.map.gz emd_14223.map.gz | 9.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14223-v30.xml emd-14223-v30.xml emd-14223.xml emd-14223.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14223_fsc.xml emd_14223_fsc.xml | 4.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_14223.png emd_14223.png | 89.2 KB | ||
| Masks |  emd_14223_msk_1.map emd_14223_msk_1.map | 10.5 MB |  Mask map Mask map | |
| Others |  emd_14223_additional_1.map.gz emd_14223_additional_1.map.gz emd_14223_half_map_1.map.gz emd_14223_half_map_1.map.gz emd_14223_half_map_2.map.gz emd_14223_half_map_2.map.gz | 5 MB 9.7 MB 9.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14223 http://ftp.pdbj.org/pub/emdb/structures/EMD-14223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14223 | HTTPS FTP |
-Related structure data
| Related structure data |  7r0jMC  7r0cC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14223.map.gz / Format: CCP4 / Size: 10.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14223.map.gz / Format: CCP4 / Size: 10.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.28 Å | ||||||||||||||||||||||||||||||||||||
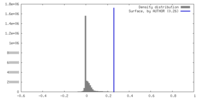
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14223_msk_1.map emd_14223_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
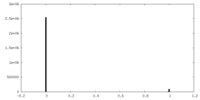
| Density Histograms |
-Additional map: #1
| File | emd_14223_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
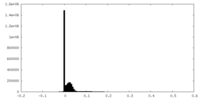
| Density Histograms |
-Half map: #2
| File | emd_14223_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
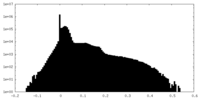
| Density Histograms |
-Half map: #1
| File | emd_14223_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of the Cterminal portion of the V2 receptor with ...
| Entire | Name: Ternary complex of the Cterminal portion of the V2 receptor with arrestin2 and ScfV30 |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of the Cterminal portion of the V2 receptor with ...
| Supramolecule | Name: Ternary complex of the Cterminal portion of the V2 receptor with arrestin2 and ScfV30 type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: V2R Cter
| Macromolecule | Name: V2R Cter / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.780248 KDa |
| Recombinant expression | Organism: Insect expression vector pBlueBacmsGCA1 (others) |
| Sequence | String: E(SEP)C(TPO)(TPO)A(SEP)(SEP)(SEP)L AKD |
-Macromolecule #2: Arrestin2
| Macromolecule | Name: Arrestin2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.164609 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MGASWSHPQF EKGGGSGGGS GGSSAWSHPQ FEKLEVLFQG PASGDKGTRV FKKASPNGKL TVYLGKRDFV DHIDLVDPVD GVVLVDPEY LKERRVYVTL TCAFRYGRED LDVLGLTFRK DLFVANVQSF PPAPEDKKPL TRLQERLIKK LGEHAYPFTF E IPPNLPCS ...String: MGASWSHPQF EKGGGSGGGS GGSSAWSHPQ FEKLEVLFQG PASGDKGTRV FKKASPNGKL TVYLGKRDFV DHIDLVDPVD GVVLVDPEY LKERRVYVTL TCAFRYGRED LDVLGLTFRK DLFVANVQSF PPAPEDKKPL TRLQERLIKK LGEHAYPFTF E IPPNLPCS VTLQPGPEDT GKACGVDYEV KAFCAENLEE KIHKRNSVRL VIRKVQYAPE RPGPQPTAET TRQFLMSDKP LH LEASLDK EIYYHGEPIS VNVHVTNNTN KTVKKIKISV RQYADICLFN TAQYKCPVAM EEADDTVAPS STFCKVYTLT PFL ANNREK RGLALDGKLK HEDTNLASST LLREGANREI LGIIVSYKVK VKLVVSRGGL LGDLASSDVA VELPFTLMHP KPKE EPPHR EVPENETPVD TNLIELDTN |
-Macromolecule #3: ScFv30
| Macromolecule | Name: ScFv30 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 30.070027 KDa |
| Recombinant expression | Organism: Insect expression vector pBlueBacmsGCA1 (others) |
| Sequence | String: AMGDIQMTQS PSSLSASVGD RVTITCRASQ SVSSAVAWYQ QKPGKAPKLL IYSASSLYSG VPSRFSGSRS GTDFTLTISS LQPEDFATY YCQQYKYVPV TFGCGTKVEI KGTTAASGSS GGSSSGAEVQ LVESGGGLVQ PGGSLRLSCA ASGFNVYSSS I HWVRQAPG ...String: AMGDIQMTQS PSSLSASVGD RVTITCRASQ SVSSAVAWYQ QKPGKAPKLL IYSASSLYSG VPSRFSGSRS GTDFTLTISS LQPEDFATY YCQQYKYVPV TFGCGTKVEI KGTTAASGSS GGSSSGAEVQ LVESGGGLVQ PGGSLRLSCA ASGFNVYSSS I HWVRQAPG KCLEWVASIS SYYGYTYYAD SVKGRFTISA DTSKNTAYLQ MNSLRAEDTA VYYCARSRQF WYSGLDYWGQ GT LVTVSSA AADDDDKAGW SHPQFEKGGG SGGGSGGGSW SHPQFEK |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 130000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 14080 / Average exposure time: 1.0 sec. / Average electron dose: 52.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)