[English] 日本語
 Yorodumi
Yorodumi- EMDB-10417: GroEL map obtained using the Preassis method for grid preparation -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10417 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GroEL map obtained using the Preassis method for grid preparation | |||||||||
 Map data Map data | GroEL map obtained using the Preassis method for sample preparation | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationGroEL-GroES complex /  chaperonin ATPase / chaperonin ATPase /  virion assembly / chaperone cofactor-dependent protein refolding / virion assembly / chaperone cofactor-dependent protein refolding /  isomerase activity / ATP-dependent protein folding chaperone / response to radiation / unfolded protein binding / isomerase activity / ATP-dependent protein folding chaperone / response to radiation / unfolded protein binding /  protein folding / response to heat ...GroEL-GroES complex / protein folding / response to heat ...GroEL-GroES complex /  chaperonin ATPase / chaperonin ATPase /  virion assembly / chaperone cofactor-dependent protein refolding / virion assembly / chaperone cofactor-dependent protein refolding /  isomerase activity / ATP-dependent protein folding chaperone / response to radiation / unfolded protein binding / isomerase activity / ATP-dependent protein folding chaperone / response to radiation / unfolded protein binding /  protein folding / response to heat / protein refolding / magnesium ion binding / protein folding / response to heat / protein refolding / magnesium ion binding /  ATP hydrolysis activity / ATP hydrolysis activity /  ATP binding / ATP binding /  membrane / identical protein binding / membrane / identical protein binding /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Escherichia coli (E. coli) Escherichia coli (E. coli) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.4 Å cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Zhao J / Xu H / Carroni M / Zou X | |||||||||
| Funding support |  Sweden, 2 items Sweden, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: A simple pressure-assisted method for MicroED specimen preparation. Authors: Jingjing Zhao / Hongyi Xu / Hugo Lebrette / Marta Carroni / Helena Taberman / Martin Högbom / Xiaodong Zou /   Abstract: Micro-crystal electron diffraction (MicroED) has shown great potential for structure determination of macromolecular crystals too small for X-ray diffraction. However, specimen preparation remains a ...Micro-crystal electron diffraction (MicroED) has shown great potential for structure determination of macromolecular crystals too small for X-ray diffraction. However, specimen preparation remains a major bottleneck. Here, we report a simple method for preparing MicroED specimens, named Preassis, in which excess liquid is removed through an EM grid with the assistance of pressure. We show the ice thicknesses can be controlled by tuning the pressure in combination with EM grids with appropriate carbon hole sizes. Importantly, Preassis can handle a wide range of protein crystals grown in various buffer conditions including those with high viscosity, as well as samples with low crystal concentrations. Preassis is a simple and universal method for MicroED specimen preparation, and will significantly broaden the applications of MicroED. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10417.map.gz emd_10417.map.gz | 25.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10417-v30.xml emd-10417-v30.xml emd-10417.xml emd-10417.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10417_fsc.xml emd_10417_fsc.xml | 12.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_10417.png emd_10417.png | 89 KB | ||
| Masks |  emd_10417_msk_1.map emd_10417_msk_1.map | 163.6 MB |  Mask map Mask map | |
| Others |  emd_10417_additional_1.map.gz emd_10417_additional_1.map.gz emd_10417_half_map_1.map.gz emd_10417_half_map_1.map.gz emd_10417_half_map_2.map.gz emd_10417_half_map_2.map.gz | 153.2 MB 129.3 MB 129.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10417 http://ftp.pdbj.org/pub/emdb/structures/EMD-10417 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10417 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10417 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10417.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10417.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GroEL map obtained using the Preassis method for sample preparation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10417_msk_1.map emd_10417_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
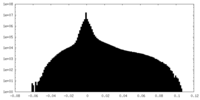
| Density Histograms |
-Additional map: postprocessed unmasked GroEL map obtained using the Preassis...
| File | emd_10417_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | postprocessed unmasked GroEL map obtained using the Preassis method for sample preparation | ||||||||||||
| Projections & Slices |
| ||||||||||||
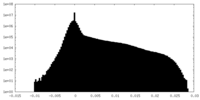
| Density Histograms |
-Half map: Half map 2 GroEL map obtained using the...
| File | emd_10417_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 GroEL map obtained using the Preassis method for sample preparation | ||||||||||||
| Projections & Slices |
| ||||||||||||
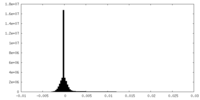
| Density Histograms |
-Half map: Half map 1 GroEL map obtained using the...
| File | emd_10417_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 GroEL map obtained using the Preassis method for sample preparation | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GroEL
| Entire | Name: GroEL |
|---|---|
| Components |
|
-Supramolecule #1: GroEL
| Supramolecule | Name: GroEL / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Details | Specimen prepared using the novel freezing system Preassis |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z
Z Y
Y X
X