+ Open data
Open data
- Basic information
Basic information
| Entry | 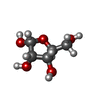 Database: PDB chemical components / ID: Z6J Database: PDB chemical components / ID: Z6J | ||
|---|---|---|---|
| Name | Name: | Wikipedia |  Wikipedia - Ribose: Wikipedia - Ribose: Ribose is a simple sugar and carbohydrate with molecular formula C5H10O5 and the linear-form composition H−(C=O)−(CHOH)4−H. The naturally-occurring form, d-ribose, is a component of the ribonucleotides from which RNA is built, and so this compound is necessary for coding, decoding, regulation and expression of genes... |
-Chemical information
| Composition |  | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Others | Type: L-saccharide, alpha linking / PDB classification: ATOMS / Three letter code: Z6J / Ideal coordinates details: Corina / Model coordinates PDB-ID: 2VGD | ||||||||||||||||
| History |
| ||||||||||||||||
 External links External links |  CompTox / CompTox /  UniChem / UniChem /  ChEBI / ChEBI /  Metabolights / Metabolights /  PubChem / PubChem /  PubChem_TPharma / PubChem_TPharma /  SureChEMBL / SureChEMBL /  ZINC / ZINC /  ChemSpider / ChemSpider /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 12.01 | | CACTVS 3.370 | OpenEye OEToolkits 1.7.6 | |
|---|
-SMILES CANONICAL
| CACTVS 3.370 | | OpenEye OEToolkits 1.7.6 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 12.01 | | OpenEye OEToolkits 1.7.6 | ( | |
|---|
-CONDENSED IUPAC CARBOHYDRATE SYMBOL
| GMML 1.0 |
|---|
-COMMON NAME
| GMML 1.0 |
|---|
-IUPAC CARBOHYDRATE SYMBOL
| PDB-CARE 1.0 |
|---|
-SNFG CARBOHYDRATE SYMBOL
| GMML 1.0 |
|---|
-PDB entries
Showing all 3 items

PDB-4q0p: 
Crystal structure of Acinetobacter sp. DL28 L-ribose isomerase in complex with L-ribose
PDB-4q0u: 
Crystal structure of Acinetobacter sp. DL28 L-ribose isomerase mutant E204Q in complex with L-ribose

PDB-4qee: 
Room temperature X-ray structure of D-xylose isomerase in complex with two Ni2+ ions and L-ribose
 Movie
Movie Controller
Controller




