+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | FZD6 Gs complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | FZD6 /  Complex. / Complex. /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationembryonic nail plate morphogenesis / establishment of body hair planar orientation /  cell proliferation in midbrain / Signaling by RNF43 mutants / midbrain morphogenesis / cell proliferation in midbrain / Signaling by RNF43 mutants / midbrain morphogenesis /  Wnt receptor activity / non-canonical Wnt signaling pathway / sensory perception of chemical stimulus / Wnt-protein binding / apicolateral plasma membrane ...embryonic nail plate morphogenesis / establishment of body hair planar orientation / Wnt receptor activity / non-canonical Wnt signaling pathway / sensory perception of chemical stimulus / Wnt-protein binding / apicolateral plasma membrane ...embryonic nail plate morphogenesis / establishment of body hair planar orientation /  cell proliferation in midbrain / Signaling by RNF43 mutants / midbrain morphogenesis / cell proliferation in midbrain / Signaling by RNF43 mutants / midbrain morphogenesis /  Wnt receptor activity / non-canonical Wnt signaling pathway / sensory perception of chemical stimulus / Wnt-protein binding / apicolateral plasma membrane / mu-type opioid receptor binding / Wnt receptor activity / non-canonical Wnt signaling pathway / sensory perception of chemical stimulus / Wnt-protein binding / apicolateral plasma membrane / mu-type opioid receptor binding /  corticotropin-releasing hormone receptor 1 binding / PCP/CE pathway / Class B/2 (Secretin family receptors) / corticotropin-releasing hormone receptor 1 binding / PCP/CE pathway / Class B/2 (Secretin family receptors) /  Wnt signaling pathway, planar cell polarity pathway / Wnt signaling pathway, planar cell polarity pathway /  beta-2 adrenergic receptor binding / inner ear morphogenesis / PKA activation in glucagon signalling / developmental growth / hair follicle development / beta-2 adrenergic receptor binding / inner ear morphogenesis / PKA activation in glucagon signalling / developmental growth / hair follicle development /  D1 dopamine receptor binding / canonical Wnt signaling pathway / Hedgehog 'off' state / Regulation of FZD by ubiquitination / D1 dopamine receptor binding / canonical Wnt signaling pathway / Hedgehog 'off' state / Regulation of FZD by ubiquitination /  insulin-like growth factor receptor binding / insulin-like growth factor receptor binding /  ionotropic glutamate receptor binding / adenylate cyclase activator activity / neural tube closure / G protein-coupled receptor activity / G-protein beta/gamma-subunit complex binding / Olfactory Signaling Pathway / Activation of the phototransduction cascade / ionotropic glutamate receptor binding / adenylate cyclase activator activity / neural tube closure / G protein-coupled receptor activity / G-protein beta/gamma-subunit complex binding / Olfactory Signaling Pathway / Activation of the phototransduction cascade /  bone development / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / cytoplasmic vesicle membrane / negative regulation of canonical Wnt signaling pathway / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / adenylate cyclase-activating G protein-coupled receptor signaling pathway / negative regulation of DNA-binding transcription factor activity / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / Sensory perception of sweet, bitter, and umami (glutamate) taste / bone development / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / cytoplasmic vesicle membrane / negative regulation of canonical Wnt signaling pathway / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / adenylate cyclase-activating G protein-coupled receptor signaling pathway / negative regulation of DNA-binding transcription factor activity / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / Sensory perception of sweet, bitter, and umami (glutamate) taste /  platelet activation / photoreceptor disc membrane / platelet activation / photoreceptor disc membrane /  platelet aggregation / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / adenylate cyclase-activating dopamine receptor signaling pathway / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / G-protein beta-subunit binding / Inactivation, recovery and regulation of the phototransduction cascade / platelet aggregation / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / adenylate cyclase-activating dopamine receptor signaling pathway / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / G-protein beta-subunit binding / Inactivation, recovery and regulation of the phototransduction cascade /  heterotrimeric G-protein complex / G alpha (12/13) signalling events / heterotrimeric G-protein complex / G alpha (12/13) signalling events /  extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye /  GTPase binding / phospholipase C-activating G protein-coupled receptor signaling pathway / Ca2+ pathway / positive regulation of cold-induced thermogenesis / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / G alpha (q) signalling events / cell population proliferation / Ras protein signal transduction / Extra-nuclear estrogen signaling / apical plasma membrane / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase binding / phospholipase C-activating G protein-coupled receptor signaling pathway / Ca2+ pathway / positive regulation of cold-induced thermogenesis / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / G alpha (q) signalling events / cell population proliferation / Ras protein signal transduction / Extra-nuclear estrogen signaling / apical plasma membrane / G protein-coupled receptor signaling pathway / lysosomal membrane /  GTPase activity / GTPase activity /  synapse / synapse /  ubiquitin protein ligase binding / protein-containing complex binding / endoplasmic reticulum membrane / GTP binding / ubiquitin protein ligase binding / protein-containing complex binding / endoplasmic reticulum membrane / GTP binding /  cell surface / cell surface /  signal transduction / extracellular exosome / signal transduction / extracellular exosome /  membrane membraneSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
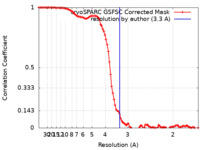
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.3 Å cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Xu F / Zhang Z | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2024 Journal: Cell Discov / Year: 2024Title: A framework for Frizzled-G protein coupling and implications to the PCP signaling pathways. Authors: Zhibin Zhang / Xi Lin / Ling Wei / Yiran Wu / Lu Xu / Lijie Wu / Xiaohu Wei / Suwen Zhao / Xiangjia Zhu / Fei Xu /  Abstract: The ten Frizzled receptors (FZDs) are essential in Wnt signaling and play important roles in embryonic development and tumorigenesis. Among these, FZD6 is closely associated with lens development. ...The ten Frizzled receptors (FZDs) are essential in Wnt signaling and play important roles in embryonic development and tumorigenesis. Among these, FZD6 is closely associated with lens development. Understanding FZD activation mechanism is key to unlock these emerging targets. Here we present the cryo-EM structures of FZD6 and FZD3 which are known to relay non-canonical planar cell polarity (PCP) signaling pathways as well as FZD1 in their G protein-coupled states and in the apo inactive states, respectively. Comparison of the three inactive/active pairs unveiled a shared activation framework among all ten FZDs. Mutagenesis along with imaging and functional analysis on the human lens epithelial tissues suggested potential crosstalk between the G-protein coupling of FZD6 and the PCP signaling pathways. Together, this study provides an integrated understanding of FZD structure and function, and lays the foundation for developing therapeutic modulators to activate or inhibit FZD signaling for a range of disorders including cancers and cataracts. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36261.map.gz emd_36261.map.gz | 80.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36261-v30.xml emd-36261-v30.xml emd-36261.xml emd-36261.xml | 20.4 KB 20.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36261_fsc.xml emd_36261_fsc.xml | 10.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_36261.png emd_36261.png | 44.7 KB | ||
| Filedesc metadata |  emd-36261.cif.gz emd-36261.cif.gz | 6.3 KB | ||
| Others |  emd_36261_additional_1.map.gz emd_36261_additional_1.map.gz emd_36261_half_map_1.map.gz emd_36261_half_map_1.map.gz emd_36261_half_map_2.map.gz emd_36261_half_map_2.map.gz | 45.1 MB 84.7 MB 84.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36261 http://ftp.pdbj.org/pub/emdb/structures/EMD-36261 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36261 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36261 | HTTPS FTP |
-Related structure data
| Related structure data |  8jhbMC  8j9nC  8j9oC  8jh7C  8jhcC  8jhiC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36261.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36261.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_36261_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36261_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36261_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : FZD6-Gs complex with Nb35.
| Entire | Name: FZD6-Gs complex with Nb35. |
|---|---|
| Components |
|
-Supramolecule #1: FZD6-Gs complex with Nb35.
| Supramolecule | Name: FZD6-Gs complex with Nb35. / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas type: protein_or_peptide / ID: 1 / Details: minGas / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 31.117201 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: HHHHHHENLY FQGSKTEDKR AQKRAEKKRS KLIDKQLQDE KMGYMCTHRL LLLGADNSGK STIVKQMRIL HGGSGGSGGT SGIFETKFQ VDKVNFHMFD VGGQRDERRK WIQCFNDVTA IIFVVDSSDY GSGGSGAGSA NRLQEALNLF KSIWNNRWLR T ISVILFLN ...String: HHHHHHENLY FQGSKTEDKR AQKRAEKKRS KLIDKQLQDE KMGYMCTHRL LLLGADNSGK STIVKQMRIL HGGSGGSGGT SGIFETKFQ VDKVNFHMFD VGGQRDERRK WIQCFNDVTA IIFVVDSSDY GSGGSGAGSA NRLQEALNLF KSIWNNRWLR T ISVILFLN KQDLLAEKVL AGKSKIEDYF PEFARYTTPE DATPEPGEDP RVTRAKYFIR DEFLRISTAS GDGRHYCYPH FT CAVDTEN ARRIFNDCRD IIQRMHLRQY ELL UniProtKB: Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas, Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas, Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.41693 KDa |
| Recombinant expression | Organism:   Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL ...String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD INAICFFPNG NA FATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGV TDDGMA VATGSWDSFL KIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.417766 KDa |
| Recombinant expression | Organism:   Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MHHHHHHNTA SIAQARKLVE QLKMEANIDR IKVSKAAADL MAYCEAHAKE DPLLTPVPAS ENPFREKKFF CAIL UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Nb35
| Macromolecule | Name: Nb35 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.845516 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MQVQLQESGG GLVQPGGSLR LSCAASGFTF SNYKMNWVRQ APGKGLEWVS DISQSGASIS YTGSVKGRFT ISRDNAKNTL YLQMNSLKP EDTAVYYCAR CPAPFTRDCF DVTSTTYAYR GQGTQVTVSS HHHHHH |
-Macromolecule #5: Frizzled-6
| Macromolecule | Name: Frizzled-6 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 61.912398 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDKHHHHHH HSLFTCEPIT VPRCMKMAYN MTFFPNLMGH YDQSIAAVEM EHFLPLANLE CSPNIETFL CKAFVPTCIE QIHVVPPCRK LCEKVYSDCK KLIDTFGIRW PEELECDRLQ YCDETVPVTF DPHTEFLGPQ K KTEQVQRD ...String: MKTIIALSYI FCLVFADYKD DDDKHHHHHH HSLFTCEPIT VPRCMKMAYN MTFFPNLMGH YDQSIAAVEM EHFLPLANLE CSPNIETFL CKAFVPTCIE QIHVVPPCRK LCEKVYSDCK KLIDTFGIRW PEELECDRLQ YCDETVPVTF DPHTEFLGPQ K KTEQVQRD IGFWCPRHLK TSGGQGYKFL GIDQCAPPCP NMYFKSDELE FAKSFIGTVS IFCLCATLFT FLTFLIDVRR FR YPERPII YYSVCYSIVS LMYFIGFLLG DSTACNKADE KLELGDTVVL GSQNKACTVL FMLLYFFTMA GTVWWVILTI TWF LAAGRK WSCEAIEQKA VWFHAVAWGT PGFLTVMLLA MNKVEGDNIS GVCFVGLYDL DASRYFVLLP LCLCVFVGLS LLLA GIISL NHVRQVIQHD GRNQEKLKKF MIRIGVFSGL YLVPLVTLLG CYVYEQVNRI TWEITWVSDH CRQYHIPCPY QAKAK ARPE LALFMIKYLM TLIVGISAVF WVGSKKTCTE WAGFFKRNRK RDPISESRRV LQE UniProtKB:  Frizzled-6 Frizzled-6 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: DIFFRACTION / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 20.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






























 Z
Z Y
Y X
X