+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10584 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PKM2 in complex with Compound 10 | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information programmed cell death / programmed cell death /  pyruvate kinase / pyruvate kinase /  pyruvate kinase activity / positive regulation of cytoplasmic translation / histone H3T11 kinase activity / canonical glycolysis / pyruvate kinase activity / positive regulation of cytoplasmic translation / histone H3T11 kinase activity / canonical glycolysis /  Glycolysis / positive regulation of sprouting angiogenesis / Glycolysis / positive regulation of sprouting angiogenesis /  potassium ion binding / potassium ion binding /  rough endoplasmic reticulum ... rough endoplasmic reticulum ... programmed cell death / programmed cell death /  pyruvate kinase / pyruvate kinase /  pyruvate kinase activity / positive regulation of cytoplasmic translation / histone H3T11 kinase activity / canonical glycolysis / pyruvate kinase activity / positive regulation of cytoplasmic translation / histone H3T11 kinase activity / canonical glycolysis /  Glycolysis / positive regulation of sprouting angiogenesis / Glycolysis / positive regulation of sprouting angiogenesis /  potassium ion binding / potassium ion binding /  rough endoplasmic reticulum / glycolytic process / rough endoplasmic reticulum / glycolytic process /  non-specific protein-tyrosine kinase / non-specific protein-tyrosine kinase /  cilium / cellular response to insulin stimulus / cilium / cellular response to insulin stimulus /  extracellular vesicle / MHC class II protein complex binding / extracellular vesicle / MHC class II protein complex binding /  protein tyrosine kinase activity / collagen-containing extracellular matrix / secretory granule lumen / vesicle / ficolin-1-rich granule lumen / protein tyrosine kinase activity / collagen-containing extracellular matrix / secretory granule lumen / vesicle / ficolin-1-rich granule lumen /  transcription coactivator activity / transcription coactivator activity /  non-specific serine/threonine protein kinase / non-specific serine/threonine protein kinase /  cadherin binding / cadherin binding /  phosphorylation / phosphorylation /  mRNA binding / Neutrophil degranulation / magnesium ion binding / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / mRNA binding / Neutrophil degranulation / magnesium ion binding / protein homodimerization activity / positive regulation of transcription by RNA polymerase II /  mitochondrion / mitochondrion /  RNA binding / extracellular exosome / extracellular region / RNA binding / extracellular exosome / extracellular region /  ATP binding / ATP binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.7 Å cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Saur M / Hartshorn MJ / Dong J / Reeks J / Bunkoczi G / Jhoti H / Williams PA | |||||||||
 Citation Citation |  Journal: Drug Discov Today / Year: 2020 Journal: Drug Discov Today / Year: 2020Title: Fragment-based drug discovery using cryo-EM. Authors: Michael Saur / Michael J Hartshorn / Jing Dong / Judith Reeks / Gabor Bunkoczi / Harren Jhoti / Pamela A Williams /  Abstract: Recent advances in electron cryo-microscopy (cryo-EM) structure determination have pushed the resolutions obtainable by the method into the range widely considered to be of utility for drug discovery. ...Recent advances in electron cryo-microscopy (cryo-EM) structure determination have pushed the resolutions obtainable by the method into the range widely considered to be of utility for drug discovery. Here, we review the use of cryo-EM in fragment-based drug discovery (FBDD) based on in-house method development. We demonstrate not only that cryo-EM can reveal details of the molecular interactions between fragments and a protein, but also that the current reproducibility, quality, and throughput are compatible with FBDD. We exemplify this using the test system β-galactosidase (Bgal) and the oncology target pyruvate kinase 2 (PKM2). | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10584.map.gz emd_10584.map.gz | 31.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10584-v30.xml emd-10584-v30.xml emd-10584.xml emd-10584.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10584.png emd_10584.png | 153.8 KB | ||
| Others |  emd_10584_additional_1.map.gz emd_10584_additional_1.map.gz emd_10584_additional_2.map.gz emd_10584_additional_2.map.gz emd_10584_half_map_1.map.gz emd_10584_half_map_1.map.gz emd_10584_half_map_2.map.gz emd_10584_half_map_2.map.gz | 159.5 MB 3 MB 134.3 MB 134.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10584 http://ftp.pdbj.org/pub/emdb/structures/EMD-10584 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10584 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10584 | HTTPS FTP |
-Related structure data
| Related structure data |  6ttqMC  6tshC  6tskC  6tteC  6ttfC  6tthC  6ttiC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10649 (Title: PKM2 in complex with Compound 10 / Data size: 745.1 EMPIAR-10649 (Title: PKM2 in complex with Compound 10 / Data size: 745.1 Data #1: Data from EPU (movies have been converted to compressed TIF) [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10584.map.gz / Format: CCP4 / Size: 39.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10584.map.gz / Format: CCP4 / Size: 39.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Relion post-process unmasked map from halfmaps which have...
| File | emd_10584_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion post-process unmasked map from halfmaps which have not been resampled after auto-refine. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: None
| File | emd_10584_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Relion auto-refine halfmap 1
| File | emd_10584_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion auto-refine halfmap 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Relion auto-refine halfmap 2
| File | emd_10584_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion auto-refine halfmap 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Pyruvate Kinase M2
| Entire | Name: Pyruvate Kinase M2 |
|---|---|
| Components |
|
-Supramolecule #1: Pyruvate Kinase M2
| Supramolecule | Name: Pyruvate Kinase M2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Molecular weight | Theoretical: 240 KDa |
-Macromolecule #1: Pyruvate kinase PKM
| Macromolecule | Name: Pyruvate kinase PKM / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number:  pyruvate kinase pyruvate kinase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 59.83182 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MGSSHHHHHH SSGLVPRGSK PHSEAGTAFI QTQQLHAAMA DTFLEHMCRL DIDSPPITAR NTGIICTIGP ASRSVETLKE MIKSGMNVA RLNFSHGTHE YHAETIKNVR TATESFASDP ILYRPVAVAL DTKGPEIRTG LIKGSGTAEV ELKKGATLKI T LDNAYMEK ...String: MGSSHHHHHH SSGLVPRGSK PHSEAGTAFI QTQQLHAAMA DTFLEHMCRL DIDSPPITAR NTGIICTIGP ASRSVETLKE MIKSGMNVA RLNFSHGTHE YHAETIKNVR TATESFASDP ILYRPVAVAL DTKGPEIRTG LIKGSGTAEV ELKKGATLKI T LDNAYMEK CDENILWLDY KNICKVVEVG SKIYVDDGLI SLQVKQKGAD FLVTEVENGG SLGSKKGVNL PGAAVDLPAV SE KDIQDLK FGVEQDVDMV FASFIRKASD VHEVRKVLGE KGKNIKIISK IENHEGVRRF DEILEASDGI MVARGDLGIE IPA EKVFLA QKMMIGRCNR AGKPVICATQ MLESMIKKPR PTRAEGSDVA NAVLDGADCI MLSGETAKGD YPLEAVRMQH LIAR EAEAA IYHLQLFEEL RRLAPITSDP TEATAVGAVE ASFKCCSGAI IVLTKSGRSA HQVARYRPRA PIIAVTRNPQ TARQA HLYR GIFPVLCKDP VQEAWAEDVD LRVNFAMNVG KARGFFKKGD VVIVLTGWRP GSGFTNTMRV VPVP |
-Macromolecule #2: 1-propan-2-yl-3-pyridin-4-yl-urea
| Macromolecule | Name: 1-propan-2-yl-3-pyridin-4-yl-urea / type: ligand / ID: 2 / Number of copies: 4 / Formula: NXH |
|---|---|
| Molecular weight | Theoretical: 179.219 Da |
| Chemical component information | 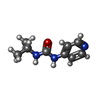 ChemComp-NXH: |
-Macromolecule #3: 1,6-di-O-phosphono-beta-D-fructofuranose
| Macromolecule | Name: 1,6-di-O-phosphono-beta-D-fructofuranose / type: ligand / ID: 3 / Number of copies: 4 / Formula: FBP |
|---|---|
| Molecular weight | Theoretical: 340.116 Da |
| Chemical component information | 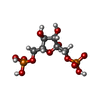 ChemComp-FBP: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 28 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.16 mg/mL |
|---|---|
| Buffer | pH: 8.2 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 572 / Average exposure time: 59.98 sec. / Average electron dose: 65.4 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 267920 |
|---|---|
| Startup model | Type of model: OTHER / Details: in-house X-ray structure |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final reconstruction | Applied symmetry - Point group: D2 (2x2 fold dihedral ) / Resolution.type: BY AUTHOR / Resolution: 2.7 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.04) / Number images used: 142864 ) / Resolution.type: BY AUTHOR / Resolution: 2.7 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.04) / Number images used: 142864 |
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Overall B value: 75.429 / Target criteria: Maximum likelihood with phases |
|---|---|
| Output model |  PDB-6ttq: |
 Movie
Movie Controller
Controller

















 Z
Z Y
Y X
X