[English] 日本語
 Yorodumi
Yorodumi- PDB-6avu: Human alpha-V beta-3 Integrin (open conformation) in complex with... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6avu | ||||||
|---|---|---|---|---|---|---|---|
| Title | Human alpha-V beta-3 Integrin (open conformation) in complex with the therapeutic antibody LM609 | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  SIGNALING PROTEIN / alpha-V beta-3 integrin / LM609 / SIGNALING PROTEIN / alpha-V beta-3 integrin / LM609 /  vitaxin / abegrin vitaxin / abegrin | ||||||
| Function / homology |  Function and homology information Function and homology informationintegrin alphav-beta6 complex / integrin alphav-beta8 complex / transforming growth factor beta production / negative regulation of entry of bacterium into host cell / integrin alphav-beta5 complex / : /  opsonin binding / integrin alphav-beta1 complex / Cross-presentation of particulate exogenous antigens (phagosomes) / opsonin binding / integrin alphav-beta1 complex / Cross-presentation of particulate exogenous antigens (phagosomes) /  extracellular matrix protein binding ...integrin alphav-beta6 complex / integrin alphav-beta8 complex / transforming growth factor beta production / negative regulation of entry of bacterium into host cell / integrin alphav-beta5 complex / : / extracellular matrix protein binding ...integrin alphav-beta6 complex / integrin alphav-beta8 complex / transforming growth factor beta production / negative regulation of entry of bacterium into host cell / integrin alphav-beta5 complex / : /  opsonin binding / integrin alphav-beta1 complex / Cross-presentation of particulate exogenous antigens (phagosomes) / opsonin binding / integrin alphav-beta1 complex / Cross-presentation of particulate exogenous antigens (phagosomes) /  extracellular matrix protein binding / tube development / regulation of serotonin uptake / positive regulation of adenylate cyclase-inhibiting opioid receptor signaling pathway / alpha9-beta1 integrin-ADAM8 complex / regulation of trophoblast cell migration / regulation of postsynaptic neurotransmitter receptor diffusion trapping / alphav-beta3 integrin-vitronectin complex / Laminin interactions / regulation of extracellular matrix organization / platelet alpha granule membrane / positive regulation of glomerular mesangial cell proliferation / integrin alphav-beta3 complex / negative regulation of lipoprotein metabolic process / alphav-beta3 integrin-PKCalpha complex / entry into host cell by a symbiont-containing vacuole / maintenance of postsynaptic specialization structure / extracellular matrix protein binding / tube development / regulation of serotonin uptake / positive regulation of adenylate cyclase-inhibiting opioid receptor signaling pathway / alpha9-beta1 integrin-ADAM8 complex / regulation of trophoblast cell migration / regulation of postsynaptic neurotransmitter receptor diffusion trapping / alphav-beta3 integrin-vitronectin complex / Laminin interactions / regulation of extracellular matrix organization / platelet alpha granule membrane / positive regulation of glomerular mesangial cell proliferation / integrin alphav-beta3 complex / negative regulation of lipoprotein metabolic process / alphav-beta3 integrin-PKCalpha complex / entry into host cell by a symbiont-containing vacuole / maintenance of postsynaptic specialization structure /  fibrinogen binding / alphav-beta3 integrin-HMGB1 complex / fibrinogen binding / alphav-beta3 integrin-HMGB1 complex /  blood coagulation, fibrin clot formation / negative regulation of lipid transport / blood coagulation, fibrin clot formation / negative regulation of lipid transport /  vascular endothelial growth factor receptor 2 binding / glycinergic synapse / negative regulation of low-density lipoprotein receptor activity / regulation of release of sequestered calcium ion into cytosol / angiogenesis involved in wound healing / Elastic fibre formation / vascular endothelial growth factor receptor 2 binding / glycinergic synapse / negative regulation of low-density lipoprotein receptor activity / regulation of release of sequestered calcium ion into cytosol / angiogenesis involved in wound healing / Elastic fibre formation /  regulation of phagocytosis / cell-substrate junction assembly / mesodermal cell differentiation / regulation of phagocytosis / cell-substrate junction assembly / mesodermal cell differentiation /  transforming growth factor beta binding / alphav-beta3 integrin-IGF-1-IGF1R complex / transforming growth factor beta binding / alphav-beta3 integrin-IGF-1-IGF1R complex /  platelet-derived growth factor receptor binding / positive regulation of small GTPase mediated signal transduction / filopodium membrane / platelet-derived growth factor receptor binding / positive regulation of small GTPase mediated signal transduction / filopodium membrane /  extracellular matrix binding / positive regulation of fibroblast migration / regulation of postsynaptic neurotransmitter receptor internalization / positive regulation of vascular endothelial growth factor receptor signaling pathway / apolipoprotein A-I-mediated signaling pathway / extracellular matrix binding / positive regulation of fibroblast migration / regulation of postsynaptic neurotransmitter receptor internalization / positive regulation of vascular endothelial growth factor receptor signaling pathway / apolipoprotein A-I-mediated signaling pathway /  regulation of bone resorption / apoptotic cell clearance / regulation of bone resorption / apoptotic cell clearance /  wound healing, spreading of epidermal cells / positive regulation of cell adhesion mediated by integrin / heterotypic cell-cell adhesion / wound healing, spreading of epidermal cells / positive regulation of cell adhesion mediated by integrin / heterotypic cell-cell adhesion /  integrin complex / Molecules associated with elastic fibres / positive regulation of intracellular signal transduction / positive regulation of cell-matrix adhesion / cellular response to insulin-like growth factor stimulus / smooth muscle cell migration / microvillus membrane / cell adhesion mediated by integrin / Syndecan interactions / negative chemotaxis / p130Cas linkage to MAPK signaling for integrins / activation of protein kinase activity / cellular response to platelet-derived growth factor stimulus / cell-substrate adhesion / integrin complex / Molecules associated with elastic fibres / positive regulation of intracellular signal transduction / positive regulation of cell-matrix adhesion / cellular response to insulin-like growth factor stimulus / smooth muscle cell migration / microvillus membrane / cell adhesion mediated by integrin / Syndecan interactions / negative chemotaxis / p130Cas linkage to MAPK signaling for integrins / activation of protein kinase activity / cellular response to platelet-derived growth factor stimulus / cell-substrate adhesion /  protein disulfide isomerase activity / positive regulation of smooth muscle cell migration / endodermal cell differentiation / positive regulation of osteoblast proliferation / TGF-beta receptor signaling activates SMADs / PECAM1 interactions / lamellipodium membrane / GRB2:SOS provides linkage to MAPK signaling for Integrins / negative regulation of macrophage derived foam cell differentiation / negative regulation of lipid storage / platelet-derived growth factor receptor signaling pathway / protein disulfide isomerase activity / positive regulation of smooth muscle cell migration / endodermal cell differentiation / positive regulation of osteoblast proliferation / TGF-beta receptor signaling activates SMADs / PECAM1 interactions / lamellipodium membrane / GRB2:SOS provides linkage to MAPK signaling for Integrins / negative regulation of macrophage derived foam cell differentiation / negative regulation of lipid storage / platelet-derived growth factor receptor signaling pathway /  fibronectin binding / positive regulation of cell adhesion / ECM proteoglycans / fibronectin binding / positive regulation of cell adhesion / ECM proteoglycans /  voltage-gated calcium channel activity / positive regulation of T cell migration / positive regulation of bone resorption / Integrin cell surface interactions / voltage-gated calcium channel activity / positive regulation of T cell migration / positive regulation of bone resorption / Integrin cell surface interactions /  vasculogenesis / vasculogenesis /  coreceptor activity / specific granule membrane / negative regulation of endothelial cell apoptotic process / positive regulation of substrate adhesion-dependent cell spreading / phagocytic vesicle / extrinsic apoptotic signaling pathway in absence of ligand / coreceptor activity / specific granule membrane / negative regulation of endothelial cell apoptotic process / positive regulation of substrate adhesion-dependent cell spreading / phagocytic vesicle / extrinsic apoptotic signaling pathway in absence of ligand /  cell adhesion molecule binding / positive regulation of endothelial cell proliferation / ERK1 and ERK2 cascade / cell adhesion molecule binding / positive regulation of endothelial cell proliferation / ERK1 and ERK2 cascade /  embryo implantation / positive regulation of endothelial cell migration / Integrin signaling / substrate adhesion-dependent cell spreading embryo implantation / positive regulation of endothelial cell migration / Integrin signaling / substrate adhesion-dependent cell spreadingSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human)  Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  negative staining / Resolution: 35 Å negative staining / Resolution: 35 Å | ||||||
 Authors Authors | Borst, A.J. / James, Z.N. / Zagotta, W.N. / Ginsberg, M. / Rey, F.A. / DiMaio, F. / Backovic, M. / Veesler, D. | ||||||
 Citation Citation |  Journal: Structure / Year: 2017 Journal: Structure / Year: 2017Title: The Therapeutic Antibody LM609 Selectively Inhibits Ligand Binding to Human αβ Integrin via Steric Hindrance. Authors: Andrew J Borst / Zachary M James / William N Zagotta / Mark Ginsberg / Felix A Rey / Frank DiMaio / Marija Backovic / David Veesler /   Abstract: The LM609 antibody specifically recognizes αβ integrin and inhibits angiogenesis, bone resorption, and viral infections in an arginine-glycine-aspartate-independent manner. LM609 entered phase II ...The LM609 antibody specifically recognizes αβ integrin and inhibits angiogenesis, bone resorption, and viral infections in an arginine-glycine-aspartate-independent manner. LM609 entered phase II clinical trials for the treatment of several cancers and was also used for αβ-targeted radioimmunotherapy. To elucidate the mechanisms of recognition and inhibition of αβ integrin, we solved the structure of the LM609 antigen-binding fragment by X-ray crystallography and determined its binding affinity for αβ. Using single-particle electron microscopy, we show that LM609 binds at the interface between the β-propeller domain of the α chain and the βI domain of the β chain, near the RGD-binding site, of all observed integrin conformational states. Integrating these data with fluorescence size-exclusion chromatography, we demonstrate that LM609 sterically hinders access of large ligands to the RGD-binding pocket, without obstructing it. This work provides a structural framework to expedite future efforts utilizing LM609 as a diagnostic or therapeutic tool. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6avu.cif.gz 6avu.cif.gz | 280.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6avu.ent.gz pdb6avu.ent.gz | 189.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6avu.json.gz 6avu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/av/6avu https://data.pdbj.org/pub/pdb/validation_reports/av/6avu ftp://data.pdbj.org/pub/pdb/validation_reports/av/6avu ftp://data.pdbj.org/pub/pdb/validation_reports/av/6avu | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7013MC 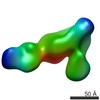 7011C 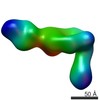 7012C  5opyC  6avqC  6avrC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein |  Integrin alpha V / Vitronectin receptor / Vitronectin receptor subunit alpha Integrin alpha V / Vitronectin receptor / Vitronectin receptor subunit alphaMass: 105894.188 Da / Num. of mol.: 1 / Fragment: UNP residues 31-987 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: ITGAV, MSK8, VNRA, VTNR / Production host: Homo sapiens (human) / Gene: ITGAV, MSK8, VNRA, VTNR / Production host:   Drosophila melanogaster (fruit fly) / References: UniProt: P06756 Drosophila melanogaster (fruit fly) / References: UniProt: P06756 |
|---|---|
| #2: Protein |  Integrin beta 3 / Platelet membrane glycoprotein IIIa / GPIIIa Integrin beta 3 / Platelet membrane glycoprotein IIIa / GPIIIaMass: 76523.125 Da / Num. of mol.: 1 / Fragment: UNP residues 27-718 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: ITGB3, GP3A / Production host: Homo sapiens (human) / Gene: ITGB3, GP3A / Production host:   Drosophila melanogaster (fruit fly) / References: UniProt: P05106 Drosophila melanogaster (fruit fly) / References: UniProt: P05106 |
| #3: Antibody | Mass: 27223.100 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Production host: Mus musculus (house mouse) / Production host:   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) |
| #4: Antibody | Mass: 23628.977 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Production host: Mus musculus (house mouse) / Production host:   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) |
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Quaternary complex of human alpha-V beta-3 integrin with the Fab LM609 Type: COMPLEX / Entity ID: all / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 0.23 MDa / Experimental value: NO |
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Source (recombinant) | Organism:   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) |
| Buffer solution | pH: 8 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied : YES / Vitrification applied : YES / Vitrification applied : NO : NO |
| EM staining | Type: NEGATIVE / Material: Uranyl formate |
| Specimen support | Grid material: COPPER / Grid mesh size: 400 divisions/in. / Grid type: C-flat 2/0.5 |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: FEI TECNAI 12 |
|---|---|
| Electron gun | Electron source : LAB6 / Accelerating voltage: 120 kV / Illumination mode: FLOOD BEAM : LAB6 / Accelerating voltage: 120 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Electron dose: 30 e/Å2 / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) |
- Processing
Processing
| EM software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
CTF correction | Type: NONE | |||||||||
3D reconstruction | Resolution: 35 Å / Resolution method: FSC 0.5 CUT-OFF / Num. of particles: 650 / Symmetry type: POINT | |||||||||
| Atomic model building | Protocol: FLEXIBLE FIT / Space: REAL |
 Movie
Movie Controller
Controller






 PDBj
PDBj













