+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30946 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Staphylothermus marinus amylopullulanase -SmApu | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | amylopullulanase /  HYDROLASE HYDROLASE | |||||||||
| Function / homology | Glycoside hydrolase family 57, N-terminal domain / Glycosyl hydrolase family 57 /  Glycoside hydrolase 38, N-terminal domain superfamily / Glycoside hydrolase/deacetylase, beta/alpha-barrel / carbohydrate metabolic process / Glycoside hydrolase 38, N-terminal domain superfamily / Glycoside hydrolase/deacetylase, beta/alpha-barrel / carbohydrate metabolic process /  hydrolase activity / hydrolase activity /  Glycoside hydrolase, family 57 Glycoside hydrolase, family 57 Function and homology information Function and homology information | |||||||||
| Biological species |   Staphylothermus marinus (archaea) / Staphylothermus marinus (archaea) /   Staphylothermus marinus (strain ATCC 43588 / DSM 3639 / JCM 9404 / F1) (archaea) Staphylothermus marinus (strain ATCC 43588 / DSM 3639 / JCM 9404 / F1) (archaea) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / cryo EM /  negative staining / Resolution: 2.9 Å negative staining / Resolution: 2.9 Å | |||||||||
 Authors Authors | Li D / Li X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Staphylothermus marinus amylopullulanase -SmApu Authors: Li D / Li X | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30946.map.gz emd_30946.map.gz | 10.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30946-v30.xml emd-30946-v30.xml emd-30946.xml emd-30946.xml | 14 KB 14 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30946_fsc.xml emd_30946_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_30946.png emd_30946.png | 90 KB | ||
| Masks |  emd_30946_msk_1.map emd_30946_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-30946.cif.gz emd-30946.cif.gz | 5.5 KB | ||
| Others |  emd_30946_half_map_1.map.gz emd_30946_half_map_1.map.gz emd_30946_half_map_2.map.gz emd_30946_half_map_2.map.gz | 80.9 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30946 http://ftp.pdbj.org/pub/emdb/structures/EMD-30946 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30946 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30946 | HTTPS FTP |
-Related structure data
| Related structure data |  7e1yMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30946.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30946.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.014 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_30946_msk_1.map emd_30946_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
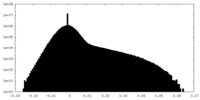
| Density Histograms |
-Half map: #2
| File | emd_30946_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
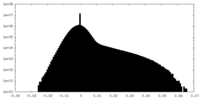
| Density Histograms |
-Half map: #1
| File | emd_30946_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
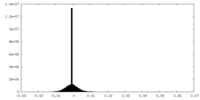
| Density Histograms |
- Sample components
Sample components
-Entire : The octamer of glycoside hydrolase family 57.
| Entire | Name: The octamer of glycoside hydrolase family 57. |
|---|---|
| Components |
|
-Supramolecule #1: The octamer of glycoside hydrolase family 57.
| Supramolecule | Name: The octamer of glycoside hydrolase family 57. / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Staphylothermus marinus (archaea) Staphylothermus marinus (archaea) |
-Macromolecule #1: Glycoside hydrolase, family 57
| Macromolecule | Name: Glycoside hydrolase, family 57 / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Staphylothermus marinus (strain ATCC 43588 / DSM 3639 / JCM 9404 / F1) (archaea) Staphylothermus marinus (strain ATCC 43588 / DSM 3639 / JCM 9404 / F1) (archaea)Strain: ATCC 43588 / DSM 3639 / JCM 9404 / F1 |
| Molecular weight | Theoretical: 75.983641 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: LEVLDKYSSL IKPKLINNIE AYMVFDKPAH KPNAEAKIYV LLNNHGSRRD IHYKIVSIDR NREVFSKRIN VDEKKFLIET ISIETPDKP GRFCYKLFID NEQIDNTCFL VGDPSSREQM YFTIVWHHHQ APNYLPDGRI HGPWAYIYVW SDLLKPYGKG P YHYHSVML ...String: LEVLDKYSSL IKPKLINNIE AYMVFDKPAH KPNAEAKIYV LLNNHGSRRD IHYKIVSIDR NREVFSKRIN VDEKKFLIET ISIETPDKP GRFCYKLFID NEQIDNTCFL VGDPSSREQM YFTIVWHHHQ APNYLPDGRI HGPWAYIYVW SDLLKPYGKG P YHYHSVML NIHPHFKATY NLSPSLLRQW QIAVEKGVEF VNGEKYDPNH EKIRLVEETL NNYREALFKG QIDVLTSIYA HT IGGFLTD VLGATNIVEE EIRYGKEVTS KIMGNNYNPQ GIWTPEMAFS MKLIPIYYDL DIKYTVLDDK FHFFHAEGNK DSQ YEPYMV IDTESKKYIT VFFRDHDLSD ILGFRNNFYS EPHAWRNAYE FALRVAEKWF DKNVKVLTIA LDGENWMSFS VNPP LTAYF LDKMIIYLET LSDNKFIKLS TLREIYNKVP ANRILTNIPT NSWLGTFRKW RGEVPQHEEY WIKTYSVYRK LLAYE EMIG GRDEFSNEAR WALWHALDSD YWWAEFWLPK IIDTWLSVAE NILNNRINKI QIIDVRPASE FYEDEKAGLV VTIRNQ LEK EIRVSFAIGG TGFSSVNNDL ETVKMNPNSS YTRIIPVKAK FIGKHKMVVS AISKGLIIDS KIIDINVKPK LLPNPRL EH HHHHH UniProtKB:  Glycoside hydrolase, family 57 Glycoside hydrolase, family 57 |
-Experimental details
-Structure determination
| Method |  negative staining, negative staining,  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.0 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Staining | Type: NEGATIVE / Material: Uranyl acetate |
| Grid | Material: COPPER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK I |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm Bright-field microscopy / Cs: 2.7 mm |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average exposure time: 8.0 sec. / Average electron dose: 64.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7e1y: |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X