+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30246 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human KCNQ4 with linopirdine | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  KCNQ / Channel / KCNQ / Channel /  Calmodulin / Calmodulin /  PIP2 / PIP2 /  linopirdine / linopirdine /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationVoltage gated Potassium channels / Sensory processing of sound by outer hair cells of the cochlea / Sensory processing of sound by inner hair cells of the cochlea / negative regulation of high voltage-gated calcium channel activity / inner ear morphogenesis / negative regulation of calcium ion export across plasma membrane / regulation of cardiac muscle cell action potential / positive regulation of ryanodine-sensitive calcium-release channel activity / regulation of cell communication by electrical coupling involved in cardiac conduction / negative regulation of peptidyl-threonine phosphorylation ...Voltage gated Potassium channels / Sensory processing of sound by outer hair cells of the cochlea / Sensory processing of sound by inner hair cells of the cochlea / negative regulation of high voltage-gated calcium channel activity / inner ear morphogenesis / negative regulation of calcium ion export across plasma membrane / regulation of cardiac muscle cell action potential / positive regulation of ryanodine-sensitive calcium-release channel activity / regulation of cell communication by electrical coupling involved in cardiac conduction / negative regulation of peptidyl-threonine phosphorylation /  voltage-gated potassium channel activity / voltage-gated potassium channel activity /  potassium channel activity / protein phosphatase activator activity / positive regulation of cyclic-nucleotide phosphodiesterase activity / positive regulation of phosphoprotein phosphatase activity / potassium channel activity / protein phosphatase activator activity / positive regulation of cyclic-nucleotide phosphodiesterase activity / positive regulation of phosphoprotein phosphatase activity /  adenylate cyclase binding / adenylate cyclase binding /  catalytic complex / detection of calcium ion / negative regulation of ryanodine-sensitive calcium-release channel activity / regulation of cardiac muscle contraction / regulation of cardiac muscle contraction by regulation of the release of sequestered calcium ion / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / regulation of calcium-mediated signaling / positive regulation of protein dephosphorylation / potassium ion transmembrane transport / catalytic complex / detection of calcium ion / negative regulation of ryanodine-sensitive calcium-release channel activity / regulation of cardiac muscle contraction / regulation of cardiac muscle contraction by regulation of the release of sequestered calcium ion / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / regulation of calcium-mediated signaling / positive regulation of protein dephosphorylation / potassium ion transmembrane transport /  voltage-gated potassium channel complex / voltage-gated potassium channel complex /  titin binding / positive regulation of protein autophosphorylation / sperm midpiece / titin binding / positive regulation of protein autophosphorylation / sperm midpiece /  calcium channel complex / substantia nigra development / adenylate cyclase activator activity / calcium channel complex / substantia nigra development / adenylate cyclase activator activity /  regulation of heart rate / regulation of heart rate /  sarcomere / protein serine/threonine kinase activator activity / sarcomere / protein serine/threonine kinase activator activity /  bioluminescence / basal plasma membrane / positive regulation of peptidyl-threonine phosphorylation / bioluminescence / basal plasma membrane / positive regulation of peptidyl-threonine phosphorylation /  regulation of cytokinesis / generation of precursor metabolites and energy / sensory perception of sound / positive regulation of protein serine/threonine kinase activity / spindle microtubule / potassium ion transport / regulation of cytokinesis / generation of precursor metabolites and energy / sensory perception of sound / positive regulation of protein serine/threonine kinase activity / spindle microtubule / potassium ion transport /  spindle pole / response to calcium ion / calcium-dependent protein binding / G2/M transition of mitotic cell cycle / spindle pole / response to calcium ion / calcium-dependent protein binding / G2/M transition of mitotic cell cycle /  myelin sheath / vesicle / transmembrane transporter binding / G protein-coupled receptor signaling pathway / myelin sheath / vesicle / transmembrane transporter binding / G protein-coupled receptor signaling pathway /  centrosome / centrosome /  calcium ion binding / calcium ion binding /  protein kinase binding / protein-containing complex / protein kinase binding / protein-containing complex /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.3 Å cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Shen H / Li T | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2021 Journal: Mol Cell / Year: 2021Title: Structural Basis for the Modulation of Human KCNQ4 by Small-Molecule Drugs. Authors: Tian Li / Kun Wu / Zhenlei Yue / Yifei Wang / Fan Zhang / Huaizong Shen /  Abstract: Among the five KCNQ channels, also known as the K7 voltage-gated potassium (K) channels, KCNQ2-KCNQ5 control neuronal excitability. Dysfunctions of KCNQ2-KCNQ5 are associated with neurological ...Among the five KCNQ channels, also known as the K7 voltage-gated potassium (K) channels, KCNQ2-KCNQ5 control neuronal excitability. Dysfunctions of KCNQ2-KCNQ5 are associated with neurological disorders such as epilepsy, deafness, and neuropathic pain. Here, we report the cryoelectron microscopy (cryo-EM) structures of human KCNQ4 and its complexes with the opener retigabine or the blocker linopirdine at overall resolutions of 2.5, 3.1, and 3.3 Å, respectively. In all structures, a phosphatidylinositol 4,5-bisphosphate (PIP) molecule inserts its head group into a cavity within each voltage-sensing domain (VSD), revealing an unobserved binding mode for PIP. Retigabine nestles in each fenestration, inducing local shifts. Instead of staying within the central pore, linopirdine resides in a cytosolic cavity underneath the inner gate. Electrophysiological analyses of various mutants corroborated the structural observations. Our studies reveal the molecular basis for the modulatory mechanism of neuronal KCNQ channels and provide a framework for structure-facilitated drug discovery targeting these important channels. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30246.map.gz emd_30246.map.gz | 115.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30246-v30.xml emd-30246-v30.xml emd-30246.xml emd-30246.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30246.png emd_30246.png | 100.9 KB | ||
| Filedesc metadata |  emd-30246.cif.gz emd-30246.cif.gz | 5.9 KB | ||
| Others |  emd_30246_additional_1.map.gz emd_30246_additional_1.map.gz | 115.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30246 http://ftp.pdbj.org/pub/emdb/structures/EMD-30246 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30246 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30246 | HTTPS FTP |
-Related structure data
| Related structure data |  7bynMC  7bylC  7bymC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30246.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30246.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: #1
| File | emd_30246_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of human KCNQ4 and linopirdine
| Entire | Name: Complex of human KCNQ4 and linopirdine |
|---|---|
| Components |
|
-Supramolecule #1: Complex of human KCNQ4 and linopirdine
| Supramolecule | Name: Complex of human KCNQ4 and linopirdine / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Green fluorescent protein,Potassium voltage-gated channel subfami...
| Macromolecule | Name: Green fluorescent protein,Potassium voltage-gated channel subfamily KQT member 4 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 109.327836 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MHHHHHHHHA ADYKDHDIDY KDDDDKSAMV SKGEELFTGV VPILVELDGD VNGHKFSVSG EGEGDATYGK LTLKFICTTG KLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYTTRA EVKFEGDTLV NRIELKGIDF K EDGNILGH ...String: MHHHHHHHHA ADYKDHDIDY KDDDDKSAMV SKGEELFTGV VPILVELDGD VNGHKFSVSG EGEGDATYGK LTLKFICTTG KLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYTTRA EVKFEGDTLV NRIELKGIDF K EDGNILGH KLEYNYNSHN VYIMADKQKN GIKVNFKIRH NIEDGSVQLA DHYQQNTPIG DGPVLLPDNH YLSTQSKLSK DP NEKRDHM VLLEFVTAAG ITLGMDELYK SGLRSGLEVL FQGPGGRMAE APPRRLGLGP PPGDAPRAEL VALTAVQSEQ GEA GGGGSP RRLGLLGSPL PPGAPLPGPG SGSGSACGQR SSAAHKRYRR LQNWVYNVLE RPRGWAFVYH VFIFLLVFSC LVLS VLSTI QEHQELANEC LLILEFVMIV VFGLEYIVRV WSAGCCCRYR GWQGRFRFAR KPFCVIDFIV FVASVAVIAA GTQGN IFAT SALRSMRFLQ ILRMVRMDRR GGTWKLLGSV VYAHSKELIT AWYIGFLVLI FASFLVYLAE KDANSDFSSY ADSLWW GTI TLTTIGYGDK TPHTWLGRVL AAGFALLGIS FFALPAGILG SGFALKVQEQ HRQKHFEKRR MPAANLIQAA WRLYSTD MS RAYLTATWYY YDSILPSFRE LALLFEHVQR ARNGGLRPLE VRRAPVPDGA PSRYPPVATC HRPGSTSFCP GESSRMGI K DRIRMGSSQR RTGPSKQHLA PPTMPTSPSS EQVGEATSPT KVQKSWSFND RTRFRASLRL KPRTSAEDAP SEEVAEEKS YQCELTVDDI MPAVKTVIRS IRILKFLVAK RKFKETLRPY DVKDVIEQYS AGHLDMLGRI KSLQTRVDQI VGRGPGDRKA REKGDKGPS DAEVVDEISM MGRVVKVEKQ VQSIEHKLDL LLGFYSRCLR SGTSASLGAV QVPLFDPDIT SDYHSPVDHE D ISVSAQTL SISRSVSTNM D UniProtKB:  Green fluorescent protein, Potassium voltage-gated channel subfamily KQT member 4 Green fluorescent protein, Potassium voltage-gated channel subfamily KQT member 4 |
-Macromolecule #2: Calmodulin-3
| Macromolecule | Name: Calmodulin-3 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 16.852545 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADQLTEEQI AEFKEAFSLF DKDGDGTITT KELGTVMRSL GQNPTEAELQ DMINEVDADG NGTIDFPEFL TMMARKMKDT DSEEEIREA FRVFDKDGNG YISAAELRHV MTNLGEKLTD EEVDEMIREA DIDGDGQVNY EEFVQMMTAK UniProtKB:  Calmodulin-3 Calmodulin-3 |
-Macromolecule #3: [(2R)-1-octadecanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tr...
| Macromolecule | Name: [(2R)-1-octadecanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phospho ryl]oxy-propan-2-yl] (8Z)-icosa-5,8,11,14-tetraenoate type: ligand / ID: 3 / Number of copies: 4 / Formula: PT5 |
|---|---|
| Molecular weight | Theoretical: 1.047088 KDa |
-Macromolecule #4: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 4 / Number of copies: 3 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Macromolecule #5: 1-phenyl-3,3-bis(pyridin-4-ylmethyl)indol-2-one
| Macromolecule | Name: 1-phenyl-3,3-bis(pyridin-4-ylmethyl)indol-2-one / type: ligand / ID: 5 / Number of copies: 1 / Formula: FCC |
|---|---|
| Molecular weight | Theoretical: 391.464 Da |
| Chemical component information | 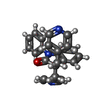 ChemComp-FCC: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.3 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 229040 |
 Movie
Movie Controller
Controller
















 Z
Z Y
Y X
X