+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30228 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Varicella-zoster virus capsid | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationT=16 icosahedral viral capsid /  viral capsid assembly / viral process / viral capsid assembly / viral process /  viral capsid / host cell nucleus / structural molecule activity / viral capsid / host cell nucleus / structural molecule activity /  DNA binding DNA bindingSimilarity search - Function | |||||||||
| Biological species | HHV-3,Human alphaherpesvirus 3 /   Human alphaherpesvirus 3 (Varicella-zoster virus) Human alphaherpesvirus 3 (Varicella-zoster virus) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.7 Å cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Wang PY / Qi JX / Liu CC / Sun JQ | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Cryo-EM structure of the varicella-zoster virus A-capsid. Authors: Junqing Sun / Congcong Liu / Ruchao Peng / Fu-Kun Zhang / Zhou Tong / Sheng Liu / Yi Shi / Zhennan Zhao / Wen-Bo Zeng / George Fu Gao / Hong-Jie Shen / Xiaoming Yang / Minhua Luo / Jianxun Qi / Peiyi Wang /  Abstract: Varicella-zoster virus (VZV), a member of the Alphaherpesvirinae subfamily, causes severe diseases in humans of all ages. The viral capsids play critical roles in herpesvirus infection, making them ...Varicella-zoster virus (VZV), a member of the Alphaherpesvirinae subfamily, causes severe diseases in humans of all ages. The viral capsids play critical roles in herpesvirus infection, making them potential antiviral targets. Here, we present the 3.7-Å-resolution structure of the VZV A-capsid and define the molecular determinants underpinning the assembly of this complicated viral machinery. Overall, the VZV capsid has a similar architecture to that of other known herpesviruses. The major capsid protein (MCP) assembles into pentons and hexons, forming extensive intra- and inter-capsomer interaction networks that are further secured by the small capsid protein (SCP) and the heterotriplex. The structure reveals a pocket beneath the floor of MCP that could potentially be targeted by antiviral inhibitors. In addition, we identified two alphaherpesvirus-specific structural features in SCP and Tri1 proteins. These observations highlight the divergence of different herpesviruses and provide an important basis for developing antiviral drugs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30228.map.gz emd_30228.map.gz | 62.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30228-v30.xml emd-30228-v30.xml emd-30228.xml emd-30228.xml | 21.8 KB 21.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30228_fsc.xml emd_30228_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_30228.png emd_30228.png | 27.9 KB | ||
| Others |  emd_30228_additional.map.gz emd_30228_additional.map.gz | 5.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30228 http://ftp.pdbj.org/pub/emdb/structures/EMD-30228 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30228 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30228 | HTTPS FTP |
-Related structure data
| Related structure data |  7bw6MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30228.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30228.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.41 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: #1
| File | emd_30228_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
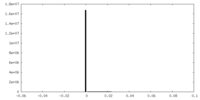
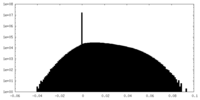
| Density Histograms |
- Sample components
Sample components
-Entire : Human alphaherpesvirus 3
| Entire | Name:   Human alphaherpesvirus 3 (Varicella-zoster virus) Human alphaherpesvirus 3 (Varicella-zoster virus) |
|---|---|
| Components |
|
-Supramolecule #1: Human alphaherpesvirus 3
| Supramolecule | Name: Human alphaherpesvirus 3 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 10335 / Sci species name: Human alphaherpesvirus 3 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|
-Macromolecule #1: Major capsid protein
| Macromolecule | Name: Major capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: HHV-3,Human alphaherpesvirus 3 |
| Molecular weight | Theoretical: 155.145359 KDa |
| Sequence | String: MTTVSCPANV ITTTESDRIA GLFNIPAGII PTGNVLSTIE VCAHRCIFDF FKQIRSDDNS LYSAQFDILL GTYCNTLNFV RFLELGLSV ACICTKFPEL AYVRDGVIQF EVQQPMIARD GPHPVDQPVH NYMVKRIHKR SLSAAFAIAS EALSLLSNTY V DGTEIDSS ...String: MTTVSCPANV ITTTESDRIA GLFNIPAGII PTGNVLSTIE VCAHRCIFDF FKQIRSDDNS LYSAQFDILL GTYCNTLNFV RFLELGLSV ACICTKFPEL AYVRDGVIQF EVQQPMIARD GPHPVDQPVH NYMVKRIHKR SLSAAFAIAS EALSLLSNTY V DGTEIDSS LRIRAIQQMA RNLRTVLDSF ERGTADQLLG VLLEKAPPLS LLSPINKFQP EGHLNRVARA ALLSDLKRRV CA DMFFMTR HAREPRLISA YLSDMVSCTQ PSVMVSRITH TNTRGRQVDG VLVTTATLKR QLLQGILQID DTAADVPVTY GEM VLQGTN LVTALVMGKA VRGMDDVARH LLDITDPNTL NIPSIPPQSN SDSTTAGLPV NARVPADLVI VGDKLVFLEA LERR VYQAT RVAYPLIGNI DITFIMPMGV FQANSMDRYT RHAGDFSTVS EQDPRQFPPQ GIFFYNKDGI LTQLTLRDAM GTICH SSLL DVEATLVALR QQHLDRQCYF GVYVAEGTED TLDVQMGRFM ETWADMMPHH PHWVNEHLTI LQFIAPSNPR LRFELN PAF DFFVAPGDVD LPGPQRPPEA MPTVNATLRI INGNIPVPLC PISFRDCRGT QLGLGRHTMT PATIKAVKDT FEDRAYP TI FYMLEAVIHG NERNFCALLR LLTQCIRGYW EQSHRVAFVN NFHMLMYITT YLGNGELPEV CINIYRDLLQ HVRALRQT I TDFTIQGEGH NGETSEALNN ILTDDTFIAP ILWDCDALIY RDEAARDRLP AIRVSGRNGY QALHFVDMAG HNFQRRDNV LIHGRPVRGD TGQGIPITPH HDREWGILSK IYYYIVIPAF SRGSCCTMGV RYDRLYPALQ AVIVPEIPAD EEAPTTPEDP RHPLHAHQL VPNSLNVYFH NAHLTVDGDA LLTLQELMGD MAERTTAILV SSAPDAGAAT ATTRNMRIYD GALYHGLIMM A YQAYDETI ATGTFFYPVP VNPLFACPEH LASLRGMTNA RRVLAKMVPP IPPFLGANHH ATIRQPVAYH VTHSKSDFNT LT YSLLGGY FKFTPISLTH QLRTGFHPGI AFTVVRQDRF ATEQLLYAER ASESYFVGQI QVHHHDAIGG VNFTLTQPRA HVD LGVGYT AVCATAALRC PLTDMGNTAQ NLFFSRGGVP MLHDNVTESL RRITASGGRL NPTEPLPIFG GLRPATSAGI ARGQ ASVCE FVAMPVSTDL QYFRTACNPR GRASGMLYMG DRDADIEAIM FDHTQSDVAY TDRATLNPWA SQKHSYGDRL YNGTY NLTG ASPIYSPCFK FFTPAEVNTN CNTLDRLLME AKAVASQSST DTEYQFKRPP GSTEMTQDPC GLFQEAYPPL CSSDAA MLR TAHAGETGAD EVHLAQYLIR DASPLRGCLP LPR |
-Macromolecule #2: Small capsomere-interacting protein
| Macromolecule | Name: Small capsomere-interacting protein / type: protein_or_peptide / ID: 2 / Number of copies: 15 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: HHV-3,Human alphaherpesvirus 3 |
| Molecular weight | Theoretical: 24.440119 KDa |
| Sequence | String: MTQPASSRVV FDPSNPTTFS VEAIAAYTPV ALIRLLNASG PLQPGHRVDI ADARSIYTVG AAASAARARA NHNANTIRRT AMFAETDPM TWLRPTVGLK RTFNPRIIRP QPPNPSMSLG ISGPTILPQK TQSADQSALQ QPAALAFSGS SPQHPPPQTT S ASVGQQQH ...String: MTQPASSRVV FDPSNPTTFS VEAIAAYTPV ALIRLLNASG PLQPGHRVDI ADARSIYTVG AAASAARARA NHNANTIRRT AMFAETDPM TWLRPTVGLK RTFNPRIIRP QPPNPSMSLG ISGPTILPQK TQSADQSALQ QPAALAFSGS SPQHPPPQTT S ASVGQQQH VVSGSSGQQP QQGAQSSTVQ PTTGSPPAAQ GVPQSTPPPT QNTPQGGKGQ TLSHTGQSGN ASRSRRV |
-Macromolecule #3: Triplex capsid protein 2
| Macromolecule | Name: Triplex capsid protein 2 / type: protein_or_peptide / ID: 3 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: HHV-3,Human alphaherpesvirus 3 |
| Molecular weight | Theoretical: 34.421914 KDa |
| Sequence | String: MAMPFEIEVL LPGELSPAET SALQKCEGKI ITFSTLRHRA SLVDIALSSY YINGAPPDTL SLLEAYRMRF AAVITRVIPG KLLAHAIGV GTPTPGLFIQ NTSPVDLCNG DYICLLPPVF GSADSIRLDS VGLEIVFPLT IPQTLMREII AKVVARAVER T AAGAQILP ...String: MAMPFEIEVL LPGELSPAET SALQKCEGKI ITFSTLRHRA SLVDIALSSY YINGAPPDTL SLLEAYRMRF AAVITRVIPG KLLAHAIGV GTPTPGLFIQ NTSPVDLCNG DYICLLPPVF GSADSIRLDS VGLEIVFPLT IPQTLMREII AKVVARAVER T AAGAQILP HEVLRGADVI CYNGRRYELE TNLQHRDGSD AAIRTLVLNL MFSINEGCLL LLALIPTLLV QGAHDGYVNL LI QTANCVR ETGQLINIPP MPRIQDGHRR FPIYETISSW ISTSSRLGDT LGTRAILRVC VFDGPSTVHP GDRTAVIQV |
-Macromolecule #4: Triplex capsid protein 1
| Macromolecule | Name: Triplex capsid protein 1 / type: protein_or_peptide / ID: 4 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: HHV-3,Human alphaherpesvirus 3 |
| Molecular weight | Theoretical: 54.02818 KDa |
| Sequence | String: MGSQPTNSHF TLNEQTLCGT NISLLGNNRF IQIGNGLHMT YAPGFFGNWS RDLTIGPRFG GLNKQPIHVP PKRTETASIQ VTPRSIVIN RMNNIQINPT SIGNPQVTIR LPLNNFKSTT QLIQQVSLTD FFRPDIEHAG SIVLILRHPS DMIGEANTLT Q AGRDPDVL ...String: MGSQPTNSHF TLNEQTLCGT NISLLGNNRF IQIGNGLHMT YAPGFFGNWS RDLTIGPRFG GLNKQPIHVP PKRTETASIQ VTPRSIVIN RMNNIQINPT SIGNPQVTIR LPLNNFKSTT QLIQQVSLTD FFRPDIEHAG SIVLILRHPS DMIGEANTLT Q AGRDPDVL LEGLRNLFNA CTAPWTVGEG GGLRAYVTSL SFIAACRAEE YTDKQAADAN RTAIVSAYGC SRMETRLIRF SE CLRAMVQ CHVFPHRFIS FFGSLLEYTI QDNLCNITAV AKGPQEAART DKTSTRRVTA NIPACVFWDV DKDLHLSADG LKH VFLVFV YTQRRQREGV RLHLALSQLN EQCFGRGIGF LLGRIRAENA AWGTEGVANT HQPYNTRALP LVQLSNDPTS PRCS IGEIT GVNWNLARQR LYQWTGDFRG LPTQLSCMYA AYTLIGTIPS ESVRYTRRME RFGGYNVPTI WLEGVVWGGT NTWNE CYY |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Homemade / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: LACEY / Support film - Film thickness: 3.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 2 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 5.116 µm / Calibrated defocus min: 0.3863 µm / Calibrated magnification: 59000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 59000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Temperature | Min: 81.0 K / Max: 93.0 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number real images: 26979 / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z
Z Y
Y X
X