+ Open data
Open data
- Basic information
Basic information
| Entry | 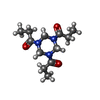 Database: PDB chemical components / ID: 29N Database: PDB chemical components / ID: 29N |
|---|---|
| Name | Name: Synonyms: 1,1',1''-(1,3,5-triazinane-1,3,5-triyl)triprop-2-en-1-one, bound form |
-Chemical information
| Composition |  | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: 29N / Ideal coordinates details: Corina / Model coordinates PDB-ID: 4MNX | ||||||||
| History |
| ||||||||
 External links External links |  UniChem / UniChem /  CompTox / CompTox /  Nikkaji / Nikkaji /  PubChem / PubChem /  SureChEMBL / SureChEMBL /  ZINC / ZINC /  ChemSpider / ChemSpider /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 12.01 | | CACTVS 3.370 | OpenEye OEToolkits 1.7.6 | |
|---|
-SMILES CANONICAL
| CACTVS 3.370 | | OpenEye OEToolkits 1.7.6 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 12.01 | | OpenEye OEToolkits 1.7.6 | |
|---|
-PDB entries
Showing all 5 items
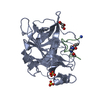
PDB-4mnx: 
Crystal structure of urokinase-type plasminogen activator (uPA) complexed with bicyclic peptide UK811

PDB-6rw2: 
Bicycle Toxin Conjugate bound to EphA2

PDB-6y8k: 
Crystal structure of CD137 in complex with the cyclic peptide BCY10916

PDB-8aaa: 
Crystal structure of SARS-CoV-2 S RBD in complex with a stapled peptide

PDB-8rtz: 
The structure of E. coli penicillin binding protein 3 (PBP3) in complex with a bicyclic peptide inhibitor
 Movie
Movie Controller
Controller



