+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12002 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the human sodium proton exchanger NHA2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationlithium:proton antiporter activity / : / positive regulation of osteoclast development / sperm principal piece / sodium:proton antiporter activity / flagellated sperm motility / regulation of insulin secretion involved in cellular response to glucose stimulus /  clathrin-dependent endocytosis / sodium ion transport / monoatomic ion transmembrane transport ...lithium:proton antiporter activity / : / positive regulation of osteoclast development / sperm principal piece / sodium:proton antiporter activity / flagellated sperm motility / regulation of insulin secretion involved in cellular response to glucose stimulus / clathrin-dependent endocytosis / sodium ion transport / monoatomic ion transmembrane transport ...lithium:proton antiporter activity / : / positive regulation of osteoclast development / sperm principal piece / sodium:proton antiporter activity / flagellated sperm motility / regulation of insulin secretion involved in cellular response to glucose stimulus /  clathrin-dependent endocytosis / sodium ion transport / monoatomic ion transmembrane transport / clathrin-dependent endocytosis / sodium ion transport / monoatomic ion transmembrane transport /  mitochondrial membrane / Stimuli-sensing channels / synaptic vesicle membrane / basolateral plasma membrane / mitochondrial membrane / Stimuli-sensing channels / synaptic vesicle membrane / basolateral plasma membrane /  mitochondrial inner membrane / endosome membrane / apical plasma membrane / identical protein binding / mitochondrial inner membrane / endosome membrane / apical plasma membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.1 Å cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Woehlert D / Kuhlbrandt W / Yildiz O | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: CryoEM structure of the human sodium proton exchanger NHA2 Authors: Woehlert D / Kuhlbrandt W / Yildiz O | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12002.map.gz emd_12002.map.gz | 77.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12002-v30.xml emd-12002-v30.xml emd-12002.xml emd-12002.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12002_fsc.xml emd_12002_fsc.xml | 9.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_12002.png emd_12002.png | 42.4 KB | ||
| Masks |  emd_12002_msk_1.map emd_12002_msk_1.map | 83.7 MB |  Mask map Mask map | |
| Others |  emd_12002_half_map_1.map.gz emd_12002_half_map_1.map.gz emd_12002_half_map_2.map.gz emd_12002_half_map_2.map.gz | 77.6 MB 77.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12002 http://ftp.pdbj.org/pub/emdb/structures/EMD-12002 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12002 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12002 | HTTPS FTP |
-Related structure data
| Related structure data |  7b4lMC  7b4mC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12002.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12002.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.073 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12002_msk_1.map emd_12002_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_12002_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_12002_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human sodium proton exchanger NHA2
| Entire | Name: Human sodium proton exchanger NHA2 |
|---|---|
| Components |
|
-Supramolecule #1: Human sodium proton exchanger NHA2
| Supramolecule | Name: Human sodium proton exchanger NHA2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) |
| Molecular weight | Experimental: 150 KDa |
-Macromolecule #1: Sodium/hydrogen exchanger 9B2
| Macromolecule | Name: Sodium/hydrogen exchanger 9B2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 57.611672 KDa |
| Recombinant expression | Organism:   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) |
| Sequence | String: MGDEDKRITY EDSEPSTGMN YTPSMHQEAQ EETVMKLKGI DANEPTEGSI LLKSSEKKLQ ETPTEANHVQ RLRQMLACPP HGLLDRVIT NVTIIVLLWA VVWSITGSEC LPGGNLFGII ILFYCAIIGG KLLGLIKLPT LPPLPSLLGM LLAGFLIRNI P VINDNVQI ...String: MGDEDKRITY EDSEPSTGMN YTPSMHQEAQ EETVMKLKGI DANEPTEGSI LLKSSEKKLQ ETPTEANHVQ RLRQMLACPP HGLLDRVIT NVTIIVLLWA VVWSITGSEC LPGGNLFGII ILFYCAIIGG KLLGLIKLPT LPPLPSLLGM LLAGFLIRNI P VINDNVQI KHKWSSSLRS IALSIILVRA GLGLDSKALK KLKGVCVRLS MGPCIVEACT SALLAHYLLG LPWQWGFILG FV LGAVSPA VVVPSMLLLQ GGGYGVEKGV PTLLMAAGSF DDILAITGFN TCLGIAFSTG STVFNVLRGV LEVVIGVATG SVL GFFIQY FPSRDQDKLV CKRTFLVLGL SVLAVFSSVH FGFPGSGGLC TLVMAFLAGM GWTSEKAEVE KIIAVAWDIF QPLL FGLIG AEVSIASLRP ETVGLCVATV GIAVLIRILT TFLMVCFAGF NLKEKIFISF AWLPKATVQA AIGSVALDTA RSHGE KQLE DYGMDVLTVA FLSILITAPI GSLLIGLLGP RLLQKVEHQN KDEEVQGETS VQV |
-Macromolecule #2: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 2 / Number of copies: 2 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #3: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 3 / Number of copies: 4 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #4: TETRADECANE
| Macromolecule | Name: TETRADECANE / type: ligand / ID: 4 / Number of copies: 6 / Formula: C14 |
|---|---|
| Molecular weight | Theoretical: 198.388 Da |
| Chemical component information |  ChemComp-C14: |
-Macromolecule #5: UNDECANE
| Macromolecule | Name: UNDECANE / type: ligand / ID: 5 / Number of copies: 10 / Formula: UND |
|---|---|
| Molecular weight | Theoretical: 156.308 Da |
| Chemical component information | 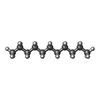 ChemComp-UND: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 26 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 4326 / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7b4l: |
 Movie
Movie Controller
Controller















 Z
Z Y
Y X
X


























