[English] 日本語
 Yorodumi
Yorodumi- EMDB-1183: Structural and functional insights into the interaction of echovi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1183 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structural and functional insights into the interaction of echoviruses and decay-accelerating factor. | |||||||||
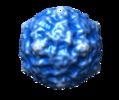
 Map data Map data | Reconstruction of Echovirus type 12 | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of lipopolysaccharide-mediated signaling pathway / negative regulation of complement activation / regulation of complement-dependent cytotoxicity / regulation of complement activation / respiratory burst / positive regulation of CD4-positive, alpha-beta T cell activation / positive regulation of CD4-positive, alpha-beta T cell proliferation / Class B/2 (Secretin family receptors) / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / ficolin-1-rich granule membrane ...regulation of lipopolysaccharide-mediated signaling pathway / negative regulation of complement activation / regulation of complement-dependent cytotoxicity / regulation of complement activation / respiratory burst / positive regulation of CD4-positive, alpha-beta T cell activation / positive regulation of CD4-positive, alpha-beta T cell proliferation / Class B/2 (Secretin family receptors) / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / ficolin-1-rich granule membrane / side of membrane / COPI-mediated anterograde transport / transport vesicle / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / endoplasmic reticulum-Golgi intermediate compartment membrane / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / complement activation, classical pathway / secretory granule membrane / T=pseudo3 icosahedral viral capsid / Regulation of Complement cascade / host cell cytoplasmic vesicle membrane / endocytosis involved in viral entry into host cell / positive regulation of T cell cytokine production / virus receptor activity / nucleoside-triphosphate phosphatase / protein complex oligomerization / monoatomic ion channel activity / positive regulation of cytosolic calcium ion concentration / DNA replication / RNA helicase activity / membrane raft / induction by virus of host autophagy / Golgi membrane / cysteine-type endopeptidase activity / RNA-directed RNA polymerase / innate immune response / viral RNA genome replication / virus-mediated perturbation of host defense response / RNA-dependent RNA polymerase activity / DNA-templated transcription / lipid binding / host cell nucleus / Neutrophil degranulation / virion attachment to host cell / structural molecule activity / cell surface / ATP hydrolysis activity / proteolysis / RNA binding / extracellular exosome / extracellular region / ATP binding / membrane / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Human echovirus 12 Human echovirus 12 | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 14.0 Å | |||||||||
 Authors Authors | Pettigrew DM / Williams DT / Kerrigan D / Evans DJ / Lea SM / Bhella D | |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2006 Journal: J Biol Chem / Year: 2006Title: Structural and functional insights into the interaction of echoviruses and decay-accelerating factor. Authors: David M Pettigrew / David T Williams / David Kerrigan / David J Evans / Susan M Lea / David Bhella /  Abstract: Many enteroviruses bind to the complement control protein decay-accelerating factor (DAF) to facilitate cell entry. We present here a structure for echovirus (EV) type 12 bound to DAF using cryo- ...Many enteroviruses bind to the complement control protein decay-accelerating factor (DAF) to facilitate cell entry. We present here a structure for echovirus (EV) type 12 bound to DAF using cryo-negative stain transmission electron microscopy and three-dimensional image reconstruction to 16-A resolution, which we interpreted using the atomic structures of EV11 and DAF. DAF binds to a hypervariable region of the capsid close to the 2-fold symmetry axes in an interaction that involves mostly the short consensus repeat 3 domain of DAF and the capsid protein VP2. A bulge in the density for the short consensus repeat 3 domain suggests that a loop at residues 174-180 rearranges to prevent steric collision between closely packed molecules at the 2-fold symmetry axes. Detailed analysis of receptor interactions between a variety of echoviruses and DAF using surface plasmon resonance and comparison of this structure (and our previous work; Bhella, D., Goodfellow, I. G., Roversi, P., Pettigrew, D., Chaudhry, Y., Evans, D. J., and Lea, S. M. (2004) J. Biol. Chem. 279, 8325-8332) with reconstructions published for EV7 bound to DAF support major differences in receptor recognition among these viruses. However, comparison of the electron density for the two virus.receptor complexes (rather than comparisons of the pseudo-atomic models derived from fitting the coordinates into these densities) suggests that the dramatic differences in interaction affinities/specificities may arise from relatively subtle structural differences rather than from large-scale repositioning of the receptor with respect to the virus surface. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1183.map.gz emd_1183.map.gz | 30.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1183-v30.xml emd-1183-v30.xml emd-1183.xml emd-1183.xml | 9.4 KB 9.4 KB | Display Display |  EMDB header EMDB header |
| Images |  1183.gif 1183.gif | 30.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1183 http://ftp.pdbj.org/pub/emdb/structures/EMD-1183 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1183 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1183 | HTTPS FTP |
-Validation report
| Summary document |  emd_1183_validation.pdf.gz emd_1183_validation.pdf.gz | 220.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1183_full_validation.pdf.gz emd_1183_full_validation.pdf.gz | 219.6 KB | Display | |
| Data in XML |  emd_1183_validation.xml.gz emd_1183_validation.xml.gz | 6.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1183 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1183 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1183 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1183 | HTTPS FTP |
-Related structure data
| Related structure data |  2c8iMC  1182C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1183.map.gz / Format: CCP4 / Size: 61.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1183.map.gz / Format: CCP4 / Size: 61.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of Echovirus type 12 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.18 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Echovirus type 12
| Entire | Name: Echovirus type 12 |
|---|---|
| Components |
|
-Supramolecule #1000: Echovirus type 12
| Supramolecule | Name: Echovirus type 12 / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Human echovirus 12
| Supramolecule | Name: Human echovirus 12 / type: virus / ID: 1 / Name.synonym: EV12 / NCBI-ID: 35293 / Sci species name: Human echovirus 12 / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No / Syn species name: EV12 |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Virus shell | Shell ID: 1 / Name: VP1-4 / Diameter: 300 Å / T number (triangulation number): 3 |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: Phosphate buffered saline |
| Staining | Type: NEGATIVE Details: 5 microlitres of sample was loaded onto a quantifoil grid and then floated on a 20 microlitre droplet of 15% Ammonium Molybdate for 10 seconds, then the grid was blotted for 2 seconds before ...Details: 5 microlitres of sample was loaded onto a quantifoil grid and then floated on a 20 microlitre droplet of 15% Ammonium Molybdate for 10 seconds, then the grid was blotted for 2 seconds before vitrification in liquid ethane. |
| Grid | Details: 400 mesh R2/2 quantifoils |
| Vitrification | Cryogen name: ETHANE / Method: Blot for two seconds, wait for two seconds. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 1200 |
|---|---|
| Temperature | Average: 100 K |
| Alignment procedure | Legacy - Astigmatism: Objective astigmatism corrected at 200,000 times mag |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON COOLSCAN / Digitization - Sampling interval: 6.35 µm / Number real images: 84 / Details: Scanned on Nikon Coolscan / Bits/pixel: 16 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Calibrated magnification: 29100 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 3.4 mm / Nominal defocus max: 2.8 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 30000 |
| Sample stage | Specimen holder: Side entry liquid-nitrogen cooled / Specimen holder model: OTHER |
- Image processing
Image processing
| CTF correction | Details: defocus pairs, each particle - CTFmix |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 14.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: PFT2 and EM3DR2 / Number images used: 1501 |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera