+Search query
-Structure paper
| Title | Cleavage-intermediate Lassa virus trimer elicits neutralizing responses, identifies neutralizing nanobodies, and reveals an apex-situated site-of-vulnerability. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 15, Issue 1, Page 285, Year 2024 |
| Publish date | Jan 4, 2024 |
 Authors Authors | Jason Gorman / Crystal Sao-Fong Cheung / Zhijian Duan / Li Ou / Maple Wang / Xuejun Chen / Cheng Cheng / Andrea Biju / Yaping Sun / Pengfei Wang / Yongping Yang / Baoshan Zhang / Jeffrey C Boyington / Tatsiana Bylund / Sam Charaf / Steven J Chen / Haijuan Du / Amy R Henry / Tracy Liu / Edward K Sarfo / Chaim A Schramm / Chen-Hsiang Shen / Tyler Stephens / I-Ting Teng / John-Paul Todd / Yaroslav Tsybovsky / Raffaello Verardi / Danyi Wang / Shuishu Wang / Zhantong Wang / Cheng-Yan Zheng / Tongqing Zhou / Daniel C Douek / John R Mascola / David D Ho / Mitchell Ho / Peter D Kwong /  |
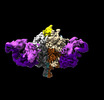
| PubMed Abstract | Lassa virus (LASV) infection is expanding outside its traditionally endemic areas in West Africa, posing a pandemic biothreat. LASV-neutralizing antibodies, moreover, have proven difficult to elicit. ...Lassa virus (LASV) infection is expanding outside its traditionally endemic areas in West Africa, posing a pandemic biothreat. LASV-neutralizing antibodies, moreover, have proven difficult to elicit. To gain insight into LASV neutralization, here we develop a prefusion-stabilized LASV glycoprotein trimer (GPC), pan it against phage libraries comprising single-domain antibodies (nanobodies) from shark and camel, and identify one, D5, which neutralizes LASV. Cryo-EM analyses reveal D5 to recognize a cleavage-dependent site-of-vulnerability at the trimer apex. The recognized site appears specific to GPC intermediates, with protomers lacking full cleavage between GP1 and GP2 subunits. Guinea pig immunizations with the prefusion-stabilized cleavage-intermediate LASV GPC, first as trimer and then as a nanoparticle, induce neutralizing responses, targeting multiple epitopes including that of D5; we identify a neutralizing antibody (GP23) from the immunized guinea pigs. Collectively, our findings define a prefusion-stabilized GPC trimer, reveal an apex-situated site-of-vulnerability, and demonstrate elicitation of LASV-neutralizing responses by a cleavage-intermediate LASV trimer. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:38177144 / PubMed:38177144 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 4.7 - 6.58 Å |
| Structure data |  EMDB-26740: Ligand-free Lassa GPC Trimer with C1 Symmetry  EMDB-26859: Ligand-free Lassa GPC Trimer with C3 Symmetry EMDB-41048, PDB-8t5c:  EMDB-41302: Lassa GPC trimer in complex with Fab GP23 |
| Chemicals |  ChemComp-NAG: |
| Source |
|
 Keywords Keywords |  VIRAL PROTEIN/IMMUNE SYSTEM / VIRAL PROTEIN/IMMUNE SYSTEM /  vaccine / GPC / GP1 / GP2 / vaccine / GPC / GP1 / GP2 /  VIRAL PROTEIN-IMMUNE SYSTEM complex VIRAL PROTEIN-IMMUNE SYSTEM complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers