+Search query
-Structure paper
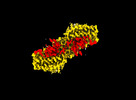
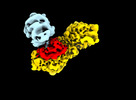
| Title | Dynamic inter-domain transformations mediate the allosteric regulation of human 5, 10-methylenetetrahydrofolate reductase. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 15, Issue 1, Page 3248, Year 2024 |
| Publish date | Apr 15, 2024 |
 Authors Authors | Linnea K M Blomgren / Melanie Huber / Sabrina R Mackinnon / Céline Bürer / Arnaud Baslé / Wyatt W Yue / D Sean Froese / Thomas J McCorvie /   |
| PubMed Abstract | 5,10-methylenetetrahydrofolate reductase (MTHFR) commits folate-derived one-carbon units to generate the methyl-donor S-adenosyl-L-methionine (SAM). Eukaryotic MTHFR appends to the well-conserved ...5,10-methylenetetrahydrofolate reductase (MTHFR) commits folate-derived one-carbon units to generate the methyl-donor S-adenosyl-L-methionine (SAM). Eukaryotic MTHFR appends to the well-conserved catalytic domain (CD) a unique regulatory domain (RD) that confers feedback inhibition by SAM. Here we determine the cryo-electron microscopy structures of human MTHFR bound to SAM and its demethylated product S-adenosyl-L-homocysteine (SAH). In the active state, with the RD bound to a single SAH, the CD is flexible and exposes its active site for catalysis. However, in the inhibited state the RD pocket is remodelled, exposing a second SAM-binding site that was previously occluded. Dual-SAM bound MTHFR demonstrates a substantially rearranged inter-domain linker that reorients the CD, inserts a loop into the active site, positions Tyr404 to bind the cofactor FAD, and blocks substrate access. Our data therefore explain the long-distance regulatory mechanism of MTHFR inhibition, underpinned by the transition between dual-SAM and single-SAH binding in response to cellular methylation status. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:38622112 / PubMed:38622112 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.8 - 3.14 Å |
| Structure data | EMDB-18298, PDB-8qa4: EMDB-18299, PDB-8qa5: EMDB-18300, PDB-8qa6: |
| Chemicals |  ChemComp-SAH:  ChemComp-HOH:  ChemComp-FAD:  ChemComp-SAM: |
| Source |
|
 Keywords Keywords |  FLAVOPROTEIN / Dis-inhibited / FLAVOPROTEIN / Dis-inhibited /  allosteric / allosteric /  folate / folate /  S-adenosylhomocysteine / Inhibited / S-Adenosylmethionin S-adenosylhomocysteine / Inhibited / S-Adenosylmethionin |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers