+Search query
-Structure paper
| Title | The SDBC is active in quenching oxidative conditions and bridges the cell envelope layers in Deinococcus radiodurans. |
|---|---|
| Journal, issue, pages | J Biol Chem, Vol. 299, Issue 1, Page 102784, Year 2023 |
| Publish date | Dec 9, 2022 |
 Authors Authors | Domenica Farci / André T Graça / Luca Iesu / Daniele de Sanctis / Dario Piano /    |
| PubMed Abstract | Deinococcus radiodurans is known for its remarkable ability to withstand harsh stressful conditions. The outermost layer of its cell envelope is a proteinaceous coat, the S-layer, essential for ...Deinococcus radiodurans is known for its remarkable ability to withstand harsh stressful conditions. The outermost layer of its cell envelope is a proteinaceous coat, the S-layer, essential for resistance to and interactions with the environment. The S-layer Deinoxanthin-binding complex (SDBC), one of the main units of the characteristic multilayered cell envelope of this bacterium, protects against environmental stressors and allows exchanges with the environment. So far, specific regions of this complex, the collar and the stalk, remained unassigned. Here, these regions are resolved by cryo-EM and locally refined. The resulting 3D map shows that the collar region of this multiprotein complex is a trimer of the protein DR_0644, a Cu-only superoxide dismutase (SOD) identified here to be efficient in quenching reactive oxygen species. The same data also showed that the stalk region consists of a coiled coil that extends into the cell envelope for ∼280 Å, reaching the inner membrane. Finally, the orientation and localization of the complex are defined by in situ cryo-electron crystallography. The structural organization of the SDBC couples fundamental UV antenna properties with the presence of a Cu-only SOD, showing here coexisting photoprotective and chemoprotective functions. These features suggests how the SDBC and similar protein complexes, might have played a primary role as evolutive templates for the origin of photoautotrophic processes by combining primary protective needs with more independent energetic strategies. |
 External links External links |  J Biol Chem / J Biol Chem /  PubMed:36502921 / PubMed:36502921 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.54 - 5.3 Å |
| Structure data | EMDB-15382: S-layer Deinoxanthin-Binding Complex, details of the stalk region  EMDB-15384: S-layer Deinoxanthin-Binding Complex, details of the N-terminal end of the stalk region  EMDB-15487: SDBC and SOD assembly, composite map  PDB-8aca:  PDB-8acq: |
| Chemicals |  ChemComp-CU: 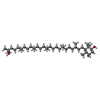 ChemComp-JPI: 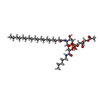 ChemComp-JQ6: 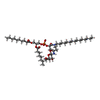 ChemComp-JPX:  ChemComp-FE: |
| Source |
|
 Keywords Keywords | METAL BINDING PROTEIN /  Superoxide Dismutase / S-layer Deinoxanthin-Binding Complex / Superoxide Dismutase / S-layer Deinoxanthin-Binding Complex /  Deinococcus radiodurans / Cell envelop / Deinococcus radiodurans / Cell envelop /  S-layer / S-layer /  STRUCTURAL PROTEIN / metal binding proteins / deinoxanthin / phosphoglycolipids / STRUCTURAL PROTEIN / metal binding proteins / deinoxanthin / phosphoglycolipids /  copper / copper /  iron iron |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers






