+Search query
-Structure paper
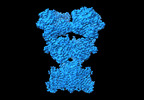
| Title | Modulation of GluA2-γ5 synaptic complex desensitization, polyamine block and antiepileptic perampanel inhibition by auxiliary subunit cornichon-2. |
|---|---|
| Journal, issue, pages | Nat Struct Mol Biol, Vol. 30, Issue 10, Page 1481-1494, Year 2023 |
| Publish date | Aug 31, 2023 |
 Authors Authors | Shanti Pal Gangwar / Laura Y Yen / Maria V Yelshanskaya / Aryeh Korman / Drew R Jones / Alexander I Sobolevsky /  |
| PubMed Abstract | Synaptic complexes of α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptors (AMPARs) with auxiliary subunits mediate most excitatory neurotransmission and can be targeted to treat ...Synaptic complexes of α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptors (AMPARs) with auxiliary subunits mediate most excitatory neurotransmission and can be targeted to treat neuropsychiatric and neurological disorders, including epilepsy. Here we present cryogenic-electron microscopy structures of rat GluA2 AMPAR complexes with inhibitory mouse γ5 and potentiating human cornichon-2 (CNIH2) auxiliary subunits. CNIH2 appears to destabilize the desensitized state of the complex by reducing the separation of the upper lobes in ligand-binding domain dimers. At the same time, CNIH2 stabilizes binding of polyamine spermidine to the selectivity filter of the closed ion channel. Nevertheless, CNIH2, and to a lesser extent γ5, attenuate polyamine block of the open channel and reduce the potency of the antiepileptic drug perampanel that inhibits the synaptic complex allosterically by binding to sites in the ion channel extracellular collar. These findings illustrate the fine-tuning of synaptic complex structure and function in an auxiliary subunit-dependent manner, which is critical for the study of brain region-specific neurotransmission and design of therapeutics for disease treatment. |
 External links External links |  Nat Struct Mol Biol / Nat Struct Mol Biol /  PubMed:37653241 / PubMed:37653241 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.72 - 3.88 Å |
| Structure data | EMDB-16052, PDB-8bhf: EMDB-40741, PDB-8ss2: EMDB-40742, PDB-8ss3: EMDB-40743, PDB-8ss4: EMDB-40744, PDB-8ss5: EMDB-40745, PDB-8ss6: EMDB-40746, PDB-8ss7: EMDB-40747, PDB-8ss8: EMDB-40748, PDB-8ss9: EMDB-40749, PDB-8ssa: EMDB-40750, PDB-8ssb: |
| Chemicals |  ChemComp-MG:  ChemComp-ZN:  ChemComp-ZK1:  ChemComp-PCW:  ChemComp-CLR:  ChemComp-AJP:  ChemComp-SPD:  ChemComp-NA:  ChemComp-6ZP:  ChemComp-HOH:  ChemComp-GLU: |
| Source |
|
 Keywords Keywords |  RIBOSOME / translational control / mRNA decay / RIBOSOME / translational control / mRNA decay /  deadenylation / ribosome stalling / deadenylation / ribosome stalling /  human CCR4-NOT / human CCR4-NOT /  MEMBRANE PROTEIN / MEMBRANE PROTEIN /  AMPA receptor / AMPA receptor /  spermidine / TARP gamma-5 / spermidine / TARP gamma-5 /  cornichon-2 / cornichon-2 /  SIGNALING PROTEIN / SIGNALING PROTEIN /  neurotransmission / neurotransmission /  Ion-channel / Ion-channel /  perampanel perampanel |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers