-検索条件
-検索結果
検索 (著者・登録者: valpuesta & j)の結果68件中、1から50件目までを表示しています

EMDB-14727: 
3D reconstruction of the cylindrical assembly of DnaJA2 by imposing D5 symmetry

EMDB-14729: 
3D reconstruction of the cylindrical assembly of DnaJA2 without symmetry imposition

EMDB-14736: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F by imposing D5 symmetry

PDB-7zhs: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F by imposing D5 symmetry

EMDB-14706: 
3D reconstruction of the cylindrical assembly of DnaJA2 delta G/F without symmetry imposition

EMDB-13588: 
Gelsolin-free CCT

EMDB-13177: 
Ser40 phosphorylated tyrosine hydroxylase

EMDB-13747: 
CCT-gelsolin complex

EMDB-13442: 
Partial structure of tyrosine hydroxylase lacking the first 35 residues in complex with dopamine.

PDB-7pim: 
Partial structure of tyrosine hydroxylase lacking the first 35 residues in complex with dopamine.

EMDB-11309: 
Partial structure of tyrosine hydroxylase in complex with dopamine showing the catalytic domain and an alpha-helix from the regulatory domain involved in dopamine binding.

PDB-6zn2: 
Partial structure of tyrosine hydroxylase in complex with dopamine showing the catalytic domain and an alpha-helix from the regulatory domain involved in dopamine binding.

EMDB-11624: 
Full-length structure of the substrate-free tyrosine hydroxylase (apo-TH).

PDB-7a2g: 
Full-length structure of the substrate-free tyrosine hydroxylase (apo-TH).

EMDB-11467: 
Atomic model of the EM-based structure of the full-length tyrosine hydroxylase in complex with dopamine (residues 40-497) in which the regulatory domain (residues 40-165) has been included only with the backbone atoms

EMDB-11587: 
Partial structure of the substrate-free tyrosine hydroxylase (apo-TH).

PDB-6zvp: 
Atomic model of the EM-based structure of the full-length tyrosine hydroxylase in complex with dopamine (residues 40-497) in which the regulatory domain (residues 40-165) has been included only with the backbone atoms

PDB-6zzu: 
Partial structure of the substrate-free tyrosine hydroxylase (apo-TH).

EMDB-10911: 
Cryo-EM structure of T7 bacteriophage DNA translocation gp15 core protein intermediate assembly

EMDB-10912: 
Cryo-EM structure of T7 bacteriophage DNA translocation gp15-gp16 core complex intermediate assembly

PDB-6ysz: 
Cryo-EM structure of T7 bacteriophage DNA translocation gp15 core protein intermediate assembly

PDB-6yt5: 
Cryo-EM structure of T7 bacteriophage DNA translocation gp15-gp16 core complex intermediate assembly

EMDB-4568: 
Cryo-EM 3D structure of CI2eng toroidal dodecameric assembly.

EMDB-0323: 
Structural insights into the ability of nucleoplasmin to assemble and chaperone histone octamers for DNA deposition

EMDB-4489: 
Human CCT:mLST8 complex

EMDB-4503: 
Human substrate-free CCT

PDB-6qb8: 
Human CCT:mLST8 complex

EMDB-4293: 
Hsp70:p53-TMGST:CHIP

EMDB-4294: 
Hsp90:p53-TMGST:CHIP

EMDB-3753: 
Negative stain 3D single particle reconstruction of the isolated surface adhesin P140/P110 complex of Mycoplasma genitalium.

EMDB-3756: 
Sub-tomogram average of the surface adhesin (NAP) complex from Mycoplasma genitalium cells by cryo-electron tomography.

EMDB-3442: 
Cryo-EM reconstructions of clathrin D6 cages

EMDB-4035: 
Cryo-EM reconstruction of clathrin D6 cages + full length Hsc70

EMDB-4036: 
Cryo-EM reconstruction of clathrin D6 cages + Hsc70 Delta C

EMDB-3232: 
The architecture of the S. pombe CCR4-NOT complex

EMDB-2447: 
Electron microscopy of the complex formed by chaperones TBCE and TBCB and alpha-tubulin

EMDB-5713: 
Electron microscopy of negatively-stained gp12 tubular protein of T7 bacteriophage

EMDB-5689: 
Cryo-electron microscopy of T7 tail complex

EMDB-5690: 
Cryo-electron microscopy of T7 tail complex formed by gp8, gp11, and gp12 proteins

PDB-3j4a: 
Structure of gp8 connector protein

PDB-3j4b: 
Structure of T7 gatekeeper protein (gp11)
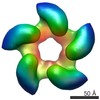
EMDB-2355: 
Negative staining three dimensional reconstruction of bacteriophage T7 large terminase

EMDB-2356: 
Negative staining three-dimensional reconstruction of bacteriophage T7 connector/terminase complex

PDB-4bij: 
Threading model of T7 large terminase

PDB-4bil: 
Threading model of the T7 large terminase within the gp8gp19 complex

EMDB-2332: 
Structural insights into the chaperone activity of Hsp40: DnaJ binds and remodels RepE

EMDB-2333: 
Structural insights into the chaperone activity of Hsp40: DnaJ binds and remodels RepE

EMDB-2334: 
Structural insights into the chaperone activity of Hsp40: DnaJ binds and remodels RepE

EMDB-2205: 
Cryo-electron microscopy reconstruction of the helical part of influenza A virus ribonucleoprotein isolated from virions.

EMDB-2206: 
Negative stained electron microscopy reconstruction of the loop of influenza A virus ribonucleoprotein isolated from virions.
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
