-Search query
-Search result
Showing 1 - 50 of 216 items for (author: zou & j)

EMDB-39212: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)
Method: single particle / : Wang CH, Chang WH

EMDB-39213: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)
Method: single particle / : Wang CH, Chang WH

EMDB-39214: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)
Method: single particle / : Wang CH, Chang WH

EMDB-39215: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)
Method: single particle / : Wang CH, Chang WH

EMDB-39217: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH5.0 (4.36A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf6: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf7: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf8: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf9: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)
Method: single particle / : Wang CH, Chang WH

EMDB-19822: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+bromosterol (DOPC, DOPE, DOPS, bromo-ergosterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ
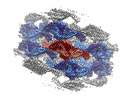

EMDB-18307: 
Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Loewith RJ, Defosses A

EMDB-18308: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture -PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol 30:20:20:30)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18309: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/-sterol (DOPC, DOPE, DOPS, PI(4,5)P2 50:20:20:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18310: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18311: 
Compact state - Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18312: 
Stretched state - Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qb7: 
Pil1 in native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Loewith RJ, Defosses A

PDB-8qb8: 
Lsp1 in native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Loewith RJ, Defosses A

PDB-8qb9: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture -PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol 30:20:20:30)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qbb: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/-sterol (DOPC, DOPE, DOPS, PI(4,5)P2 50:20:20:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qbd: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qbe: 
Compact state - Pil1 in native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qbf: 
Compact state - Pil1 dimer with lipid headgroups fitted in native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qbg: 
Stretched state - Pil1 in native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-38860: 
structure of RSF-147bp NCP complex Class 0
Method: single particle / : Zhang JL
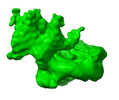

EMDB-38861: 
Structure of RSF-147bpNCP complex class 2
Method: single particle / : Zhang JL

EMDB-38865: 
RSF-38N38NCP complex Class 2
Method: single particle / : Zhang JL

EMDB-50019: 
cryoEM structure of Photosystem II averaged across S2-S3 states at 1.71 Angstrom resolution
Method: single particle / : Hussein R, Graca A, Zouni A, Messinger J, Schroder WP

PDB-9evx: 
cryoEM structure of Photosystem II averaged across S2-S3 states at 1.71 Angstrom resolution
Method: single particle / : Hussein R, Graca A, Zouni A, Messinger J, Schroder WP

EMDB-36721: 
Structure of TbAQP2 in complex with anti-trypanosomatid drug melarsoprol
Method: single particle / : Chen W, Wang C

EMDB-36722: 
Structure of the TbAQP2 in the apo conformation
Method: single particle / : Chen W, Wang C

EMDB-36723: 
Structure of TbAQP2 in complex with anti-trypanosomatid drug pentamidine
Method: single particle / : Chen W, Wang C

PDB-8jy6: 
Structure of TbAQP2 in complex with anti-trypanosomatid drug melarsoprol
Method: single particle / : Chen W, Wang C

PDB-8jy8: 
Structure of TbAQP2 in complex with anti-trypanosomatid drug pentamidine
Method: single particle / : Chen W, Wang C

EMDB-50218: 
Negative staining EM map for Mis18 core complex
Method: single particle / : Jeyaprakash AA, Medina-Pritchard B

EMDB-50219: 
Negative staining EM map for Mis18 core complex
Method: single particle / : Jeyaprakash AA, Medina-Pritchard B

EMDB-50220: 
Negative staining EM map for Mis18 core complex
Method: single particle / : Jeyaprakash AA, Medina-Pritchard B

EMDB-36907: 
Cryo-EM structure of the RC-LH core comples from Halorhodospira halochloris
Method: single particle / : Wang GL, Qi CH, Yu LJ

PDB-8k5o: 
Cryo-EM structure of the RC-LH core comples from Halorhodospira halochloris
Method: single particle / : Wang GL, Qi CH, Yu LJ
EMDB-37130: 
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form
Method: single particle / : Xie J, Wang L, Zhai G, Wu D, Lin Z, Wang M, Yan X, Gao L, Huang X, Fearns R, Chen S

EMDB-37131: 
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in monomeric form
Method: single particle / : Xie J, Wang L, Zhai G, Wu D, Lin Z, Wang M, Yan X, Gao L, Huang X, Fearns R, Chen S

PDB-8kdb: 
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form
Method: single particle / : Xie J, Wang L, Zhai G, Wu D, Lin Z, Wang M, Yan X, Gao L, Huang X, Fearns R, Chen S

PDB-8kdc: 
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in monomeric form
Method: single particle / : Xie J, Wang L, Zhai G, Wu D, Lin Z, Wang M, Yan X, Gao L, Huang X, Fearns R, Chen S

EMDB-38864: 
RSF-38N38NCP complex Class0
Method: single particle / : Zhang JL

EMDB-35727: 
Cryo-EM structure of the CRT-LESS RC-LH core complex from roseiflexus castenholzii
Method: single particle / : Wang GL, Qi CH, Yu LJ

PDB-8iun: 
Cryo-EM structure of the CRT-LESS RC-LH core complex from roseiflexus castenholzii
Method: single particle / : Wang GL, Qi CH, Yu LJ
EMDB-34711: 
Cryo-EM structure of human ZDHHC9/GCP16 complex
Method: single particle / : Wu J, Hu Q, Zhang Y, Liu S, Yang A
EMDB-34717: 
Cryo-EM structure of yeast Erf2/Erf4 complex
Method: single particle / : Wu J, Hu Q, Zhang Y, Yang A, Liu S

PDB-8hf3: 
Cryo-EM structure of human ZDHHC9/GCP16 complex
Method: single particle / : Wu J, Hu Q, Zhang Y, Liu S, Yang A
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
