-Search query
-Search result
Showing all 48 items for (author: uchiyama & a)
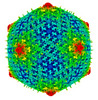

EMDB-32974: 
Capsid structure of Staphylococcus jumbo bacteriophage S6
Method: single particle / : Koibuchi W, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

PDB-7x30: 
Capsid structure of Staphylococcus jumbo bacteriophage S6
Method: single particle / : Koibuchi W, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-34741: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-11
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-34742: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-11 focused on RBD and NIV-11 interface
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

PDB-8hgl: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-11
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

PDB-8hgm: 
Structure of SARS-CoV-2 spike RBD in complex with neutralizing antibody NIV-11
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-34274: 
Bre1-nucleosome complex
Method: single particle / : Onishi S, Hamada K, Sato K, Nishizawa T, Nureki O, Ogata K, Sengoku T

EMDB-34275: 
Human nucleosome core particle (free form)
Method: single particle / : Onishi S, Sato K, Nishizawa T, Nureki O, Ogata K, Sengoku T

PDB-8gui: 
Bre1-nucleosome complex (Model I)
Method: single particle / : Onishi S, Hamada K, Sato K, Nishizawa T, Nureki O, Ogata K, Sengoku T

PDB-8guj: 
Bre1-nucleosome complex (Model II)
Method: single particle / : Onishi S, Sato K, Hamada K, Nishizawa T, Nureki O, Ogata K, Sengoku T

PDB-8guk: 
Human nucleosome core particle (free form)
Method: single particle / : Onishi S, Sato K, Nishizawa T, Nureki O, Ogata K, Sengoku T

EMDB-33820: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-8 focused on RBD and NIV-8 interface
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33821: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-8 (state 1)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33822: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-8 (state 2)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33823: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-10 focused on RBD and NIV-10 interface
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33824: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-10 (state 1)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33825: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-10 (state 2)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33826: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-10 (state 3)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33827: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-13 focused on RBD and NIV-13 interface
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33828: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-13 (state 1)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33829: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-13 (state 2)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-33830: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-13 (state 3)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

PDB-7yh6: 
Structure of SARS-CoV-2 spike RBD in complex with neutralizing antibody NIV-8
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

PDB-7yh7: 
SARS-CoV-2 spike in complex with neutralizing antibody NIV-8 (state 2)
Method: single particle / : Moriyama S, Anraku Y, Muranishi S, Adachi Y, Kuroda D, Higuchi Y, Kotaki R, Tonouchi K, Yumoto K, Suzuki T, Kita S, Fukuhara H, Kuroda Y, Yamamoto T, Onodera T, Fukushi S, Maeda K, Nakamura-Uchiyama F, Hashiguchi T, Hoshino A, Maenaka K, Takahashi Y

EMDB-32614: 
Overall structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-32616: 
Tail structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

PDB-7wmp: 
Tail structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-30778: 
Acidic stable capsid structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-30800: 
Acidic stable capsid structure of Helicobacter pylori bacteriophage KHP40
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

PDB-7dn2: 
Acidic stable capsid structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

PDB-7dou: 
Trimeric cement protein structure of Helicobacter pylori bacteriophage KHP40
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

PDB-7f2p: 
The head structure of Helicobacter pylori bacteriophage KHP40
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-0834: 
Structure of the peripheral FCPI from diatom
Method: single particle / : Nagao R, Kato K, Miyazaki N, Akita F, Shen JR

EMDB-0835: 
Structure of the PSI-FCPI supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Miyazaki N, Akita F, Shen JR

PDB-6l4t: 
Structure of the peripheral FCPI from diatom
Method: single particle / : Nagao R, Kato K, Miyazaki N, Akita F, Shen JR

PDB-6l4u: 
Structure of the PSI-FCPI supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Miyazaki N, Akita F, Shen JR

EMDB-0755: 
Recombinant Pyrococcus furiosus PbaA/PF0014 with 30 residue C-terminal truncation.
Method: single particle / : Burton-Smith RN, Song C, Murata K

EMDB-9775: 
Structure of C2S2-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Shen JR, Miyazaki N, Akita F

EMDB-9776: 
Structure of C2S1M1-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Shen JR, Miyazaki N, Akita F

EMDB-9777: 
Structure of C2S2M2-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Shen JR, Miyazaki N, Akita F

PDB-6j3y: 
Structure of C2S2-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Shen JR, Miyazaki N, Akita F

PDB-6j3z: 
Structure of C2S1M1-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Shen JR, Miyazaki N, Akita F

PDB-6j40: 
Structure of C2S2M2-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Shen JR, Miyazaki N, Akita F

EMDB-2662: 
Structure of SMG1 kinase
Method: single particle / : Melero R, Uchiyama A, Castano R, Kataoka N, Kurosawa H, Ohno S, Yamashita A, Llorca O
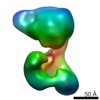
EMDB-2663: 
Structure of SMG1C complex, comprising SMG1 kinase, SMG8 and SMG9
Method: single particle / : Melero R, Uchiyama A, Castano R, Kataoka N, Kurosawa H, Ohno S, Yamashita A, Llorca O
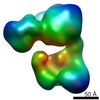
EMDB-2664: 
Structure of SMG1C-UPF1 complex, comprising SMG1 kinase, SMG8, SMG9 and UPF1
Method: single particle / : Melero R, Uchiyama A, Castano R, Kataoka N, Kurosawa H, Ohno S, Yamashita A, Llorca O

EMDB-2665: 
Structure of SMG1C-UPF2 complex, comprising SMG1 kinase, SMG8, SMG9 and UPF2
Method: single particle / : Melero R, Uchiyama A, Castano R, Kataoka N, Kurosawa H, Ohno S, Yamashita A, Llorca O

EMDB-2666: 
Structure of SMG1C-UPF1-UPF2 complex, comprising SMG1 kinase, SMG8, SMG9, UPF1 and UPF2
Method: single particle / : Melero R, Uchiyama A, Castano R, Kataoka N, Kurosawa H, Ohno S, Yamashita A, Llorca O
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
