-Search query
-Search result
Showing 1 - 50 of 87 items for (author: smith & jl)

EMDB-17767: 
Cryo electron tomography of human choriocarcinoma cells
Method: electron tomography / : Tun WM, Yee NB-Y, Ho EML, Darrow MC, Basham M

EMDB-14088: 
Icosahedral capsid of the phicrAss001 virion
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-14089: 
Portal protein assembly of the phicrAss001 virion with C12 symmetry imposed
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-14090: 
Unique vertex of the phicrAss001 virion with C5 symmetry imposed
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-14091: 
Unique vertex of the phicrAss001 virion
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-14092: 
Tail barrel assembly of the phicrAss001 virion with C12 symmetry imposed
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-14093: 
Tail muzzle assembly of the phicrAss001 virion with C6 symmetry imposed
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-14094: 
Tail assembly of the phicrAss001 virion with C6 symmetry imposed
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-14100: 
Asymmetric reconstruction of the phicrAss001 virion
Method: single particle / : Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C, Antson AA

EMDB-27203: 
IMM20190 Fab complex with SARS-CoV-2 Spike Trimer
Method: single particle / : Nikitin PA, DiMuzio JD, Dowling JP, Patel NB, Bingaman-Steele JL, Heimbach BC, Henriquez N, Nicolescu C, Polley A, Sikorski EL, Howanski RJ, Nath M, Shukla H, Scheaffer SM, Finn JP, Liang LF, Smith T, Storm N, McKay LGA, Johnson RI, Malsick LE, Honko AN, Griffiths A, Diamond MS, Sarma P, Geising DH, Morin MJ, Robinson MK

EMDB-26372: 
SARS-2 CoV 6P Mut7 in complex with Fab CC84.5
Method: single particle / : Torres JL, Ward AB

EMDB-27192: 
IMM20253 Fab complex with Trimer Spike protein of SARS-CoV-2 virus
Method: single particle / : Nikitin PA, DiMuzio JD, Dowling JP, Patel NB, Bingaman-Steele JL, Heimbach BC, Henriquez N, Nicolescu C, Polley A, Sikorski EL, Howanski RJ, Nath M, Shukla H, Scheaffer SM, Finn JP, Liang LF, Smith T, Storm N, McKay LGA, Johnson RI, Malsick LE, Honko AN, Griffiths A, Diamond MS, Sarma P, Geising DH, Morin MJ, Robinson MK

EMDB-27193: 
IMM20253 Fab complex with Spike monomer
Method: single particle / : Nikitin PA, DiMuzio JD, Dowling JP, Patel NB, Bingaman-Steele JL, Heimbach BC, Henriquez N, Nicolescu C, Polley A, Sikorski EL, Howanski RJ, Nath M, Shukla H, Scheaffer SM, Finn JP, Liang LF, Smith T, Storm N, McKay LGA, Johnson RI, Malsick LE, Honko AN, Griffiths A, Diamond MS, Sarma P, Geising DH, Morin MJ, Robinson MK

EMDB-27204: 
IMM20184 Fab complex with SARS-CoV-2 Spike Trimer
Method: single particle / : Nikitin PA, DiMuzio JD, Dowling JP, Patel NB, Bingaman-Steele JL, Heimbach BC, Henriquez N, Nicolescu C, Polley A, Sikorski EL, Howanski RJ, Nath M, Shukla H, Scheaffer SM, Finn JP, Liang LF, Smith T, Storm N, McKay LGA, Johnson RI, Malsick LE, Honko AN, Griffiths A, Diamond MS, Sarma P, Geising DH, Morin MJ, Robinson MK

EMDB-26522: 
SARS-CoV-2 6P Mut7 in complex with K398.25 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26523: 
SARS-CoV-1 in complex with K398.25 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26524: 
SARS-CoV-2 6P Mut7 in complex with K398.16 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB
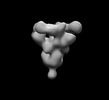
EMDB-26525: 
SARS-CoV-2 6P Mut7 in complex with K398.16 Fab (3 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26526: 
SARS-CoV-1 in complex with K398.16 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26527: 
SARS-CoV-2 6P Mut7 in complex with K288.2 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26528: 
SARS-CoV-2 6P Mut7 in complex with K398.8 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26529: 
SARS-CoV-2 6P Mut7 in complex with K398.8 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26530: 
SARS-CoV-2 6P Mut7 in complex with K398.18 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26531: 
SARS-CoV-2 6P Mut7 in complex with K398.18 Fabs (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26532: 
SARS-CoV-2 6P Mut7 in complex with K398.22 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB
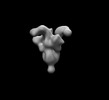
EMDB-26533: 
SARS-CoV-2 6P Mut7 in complex with K398.22 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26534: 
SARS-CoV-1 in complex with K398.8 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26535: 
SARS-CoV-1 in complex with K398.8 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26536: 
SARS-CoV-1 in complex with K398.18 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26537: 
SARS-CoV-1 in complex with K398.18 Fab (3 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26538: 
SARS-CoV-1 in complex with K288.2 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26539: 
SARS-CoV-1 in complex with K398.22 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-27081: 
IMM20184 and IMM20253 Fabs in ternary complex with SARS-CoV-2 receptor binding domain
Method: single particle / : Robinson MK, Nikitin PN

EMDB-26365: 
SARS-2 CoV 6P Mut7 in complex with Fab CC25.1
Method: single particle / : Torres JL, Ward AB
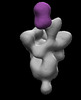
EMDB-26366: 
SARS-2 CoV 6P Mut7 in complex with Fab CC25.54
Method: single particle / : Torres JL, Ward AB

EMDB-26367: 
SARS-2 CoV 6P Mut7 + Fab CC25.52
Method: single particle / : Torres JL, Ward AB

EMDB-26368: 
SARS-2 CoV 6P Mut7 in complex with Fab CC25.56
Method: single particle / : Torres JL, Ward AB

EMDB-26369: 
SARS-2 CoV 6P Mut7 in complex with Fab CC25.36
Method: single particle / : Torres JL, Ward AB

EMDB-26370: 
SARS-2 CoV 6P Mut7 in complex with Fab CC25.3
Method: single particle / : Torres JL, Ward AB

EMDB-26371: 
SARS-2 CoV 6P Mut7 in complex with Fab CC25.11
Method: single particle / : Torres JL, Ward AB

EMDB-26373: 
SARS-2 CoV 6P Mut7 in complex with Fab CC84.2
Method: single particle / : Torres JL, Ward AB

EMDB-26374: 
SARS-2 CoV 6P Mut7 in complex with Fab CC84.10
Method: single particle / : Torres JL, Ward AB

EMDB-26375: 
SARS-2 CoV 6P Mut7 in complex with Fab CC84.24
Method: single particle / : Torres JL, Ward AB

EMDB-12590: 
Bacteriophage YerA41 head icosahedral reconstruction
Method: single particle / : Gomez-Raya-Vilanova MV, Leskinen K, Bhattacharjee A, Virta P, Rosenqvist P, Smith JLR, Bayfield OW, Homberger C, Kerrinnes T, Voge J, Pajunen MI, Skurnik M

EMDB-12311: 
Structure of Wild-Type Human Potassium Chloride Transporter KCC3 in NaCl (LMNG/CHS)
Method: single particle / : Chi G, Man H, Pike ACW, Wang D, McKinley G, Mukhopadhyay SMM, MacLean EM, Chalk R, Moreau C, Snee M, Abrusci P, Arrowsmith CH, Bountra C, Edwards AM, Marsden BD, Burgess-Brown NA, Duerr KL

EMDB-11799: 
Structure of Human Potassium Chloride Transporter KCC3 S45D/T940D/T997D in NaCl (Reference Map)
Method: single particle / : Chi G, Man H, Ebenhoch R, Reggiano G, Pike ACW, Wang D, McKinley G, Mukhopadhyay SMM, MacLean EM, Chalk R, Moreau C, Snee M, Bohstedt T, Singh NK, Abrusci P, Arrowsmith CH, Bountra C, Edwards AM, Marsden BD, Burgess-Brown NA, DiMaio F, Duerr KL, Structural Genomics Consortium (SGC)

EMDB-11800: 
Structure of Human Potassium Chloride Transporter KCC3 S45D/T940D/T997D in NaCl (Subclass)
Method: single particle / : Chi G, Man H, Ebenhoch R, Reggiano G, Pike ACW, Wang D, McKinley G, Mukhopadhyay SMM, MacLean EM, Chalk R, Moreau C, Snee M, Bohstedt T, Singh NK, Abrusci P, Arrowsmith CH, Bountra C, Edwards AM, Marsden BD, Burgess-Brown NA, DiMaio F, Duerr KL, Structural Genomics Consortium (SGC)

EMDB-11801: 
Structure of Human Potassium Chloride Transporter KCC1 in NaCl (Reference Map)
Method: single particle / : Ebenhoch R, Chi G, Man H, Wang D, McKinley G, Mukhopadhyay SMM, MacLean EM, Chalk R, Moreau C, Snee M, Bohstedt T, Liko I, Tehan BG, Almeida FG, Elkins J, Singh NK, Abrusci P, Arrowsmith CH, Tang H, Robinson CV, Bountra C, Edwards AM, Marsden BD, Burgess-Brown NA, Duerr KL, Structural Genomics Consortium (SGC)

EMDB-11802: 
Structure of Human Potassium Chloride Transporter KCC1 in NaCl (Subclass 1)
Method: single particle / : Ebenhoch R, Chi G, Man H, Wang D, McKinley G, Mukhopadhyay SMM, MacLean EM, Chalk R, Moreau C, Snee M, Bohstedt T, Singh NK, Abrusci P, Liko I, Tehan BG, Almeida FG, Arrowsmith CH, Tang H, Robinson CV, Bountra C, Edwards AM, Marsden BD, Burgess-Brown NA, Duerr KL, Structural Genomics Consortium (SGC)

EMDB-11803: 
Structure of Human Potassium Chloride Transporter KCC1 in NaCl (Subclass 2)
Method: single particle / : Ebenhoch R, Chi G, Man H, Wang D, McKinley G, Mukhopadhyay SMM, MacLean EM, Chalk R, Moreau C, Snee M, Bohstedt T, Singh NK, Abrusci P, Liko I, Tehan BG, Almeida FG, Arrowsmith CH, Tang H, Robinson CV, Bountra C, Edwards AM, Marsden BD, Burgess-Brown NA, Duerr KL, Structural Genomics Consortium (SGC)
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
