-Search query
-Search result
Showing 1 - 50 of 102 items for (author: smith & dm)

EMDB-28876: 
Thermoplasma acidophilum 20S proteasome - L81Y mutation in alpha subunit
Method: single particle / : Chuah J, Smith D

EMDB-28878: 
Thermoplasma acidophilum 20S proteasome - wild type
Method: single particle / : Chuah J, Smith D

EMDB-28906: 
Thermoplasma acidophilum 20S proteasome - wild type bound to ZYA
Method: single particle / : Chuah J, Smith D
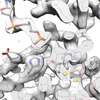
EMDB-29307: 
Structure of WT HIV-1 intasome bound to Dolutegravir
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29309: 
Structure of E138K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29312: 
Structure of E138K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29313: 
Structure of Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29315: 
Structure of E138K/G140A HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29317: 
Structure of E138K/Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29318: 
Structure of G140A/Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29319: 
Structure of E138K/G140A/Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29320: 
Structure of E138K/G140A/Q148R HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29321: 
Structure of E138K/G140S/Q148H HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

EMDB-29322: 
Structure of E138K/G140A/Q148K HIV-1 intasome with 4d bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fn7: 
Structure of WT HIV-1 intasome bound to Dolutegravir
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnd: 
Structure of E138K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fng: 
Structure of E138K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnh: 
Structure of Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnj: 
Structure of E138K/G140A HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnl: 
Structure of E138K/Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnm: 
Structure of G140A/Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnn: 
Structure of E138K/G140A/Q148K HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fno: 
Structure of E138K/G140A/Q148R HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnp: 
Structure of E138K/G140S/Q148H HIV-1 intasome with Dolutegravir bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D

PDB-8fnq: 
Structure of E138K/G140A/Q148K HIV-1 intasome with 4d bound
Method: single particle / : Shan ZL, Passos DO, Strutzenberg TS, Li M, Lyumkis D
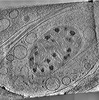
EMDB-29207: 
CryoET tomogram of mitochondria in BACHD mouse model neuron
Method: electron tomography / : Wu GH, Galaz-Montoya JG, Gu Y, Mitchell PG, Wu C, Chiu W
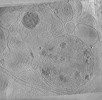
EMDB-29208: 
CryoET tomogram of BACHD mouse model neuron showing sheet aggregates
Method: electron tomography / : Wu GH, Galaz-Montoya JG, Gu Y, Mitchell PG, Wu C, Chiu W
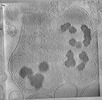
EMDB-29210: 
CryoET tomogram of purified mitochondria from HD patient iPSC-derived neuron (Q109)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
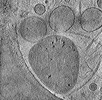
EMDB-29211: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) with PIAS1 hetKO treatment
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-26809: 
KDM2A-nucleosome structure stabilized by H3K36C-UNC8015 covalent conjugate
Method: single particle / : Spangler CJ, Skrajna A, Foley CA, Budziszewski GR, Azzam DN, James LI, Frye SV, McGinty RK

EMDB-26810: 
KDM2B-nucleosome complex stabilized by H3K36C-UNC8015 covalent conjugate
Method: single particle / : Spangler CJ, Skrajna A, Foley CA, Budziszewski GR, Azzam DN, James LI, Frye SV, McGinty RK

PDB-7uv9: 
KDM2A-nucleosome structure stabilized by H3K36C-UNC8015 covalent conjugate
Method: single particle / : Spangler CJ, Skrajna A, Foley CA, Budziszewski GR, Azzam DN, James LI, Frye SV, McGinty RK

EMDB-28668: 
CryoET tomogram of iPSC-derived control non-HD neuron (Q18)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
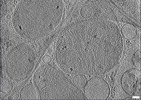
EMDB-28944: 
CryoET tomogram of iPSC-derived control non-HD neuron (Q20)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
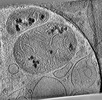
EMDB-28946: 
CryoET tomogram of HD patient iPSC-derived neuron (Q53)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
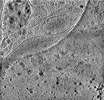
EMDB-29074: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29075: 
CryoET tomogram of HD patient iPSC-derived neuron (Q77)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29076: 
CryoET tomogram of HD patient iPSC-derived neuron (Q109)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29079: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) showing sheet aggregate
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29080: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) with PIAS1 hetKO treatment
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
EMDB-29081: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) with GRSF1 KD treatment
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29083: 
CryoET tomogram of mitochondria in WT mouse neuron
Method: electron tomography / : Wu GH, Galaz-Montoya JG, Gu Y, Mitchell PG, Wu C, Chiu W

EMDB-29084: 
CryoET tomogram of mitochondria in BACHD-dN17 mouse model neuron
Method: electron tomography / : Wu GH, Galaz-Montoya JG, Gu Y, Mitchell PG, Wu C, Chiu W

EMDB-26862: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD
Method: helical / : Li S, Nanson JD, Manik MK, Gu W, Landsberg MJ, Ve T, Kobe B

PDB-7uxu: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD
Method: helical / : Li S, Nanson JD, Manik MK, Gu W, Landsberg MJ, Ve T, Kobe B

EMDB-26379: 
Structure of PA28gamma bound to the human 20S proteasome with C7 symmetry.
Method: single particle / : Smith DM, Thomas T

EMDB-26372: 
SARS-2 CoV 6P Mut7 in complex with Fab CC84.5
Method: single particle / : Torres JL, Ward AB

EMDB-26522: 
SARS-CoV-2 6P Mut7 in complex with K398.25 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26523: 
SARS-CoV-1 in complex with K398.25 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26524: 
SARS-CoV-2 6P Mut7 in complex with K398.16 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
