-Search query
-Search result
Showing 1 - 50 of 73 items for (author: moran & a)

EMDB-41144: 
Cryo-EM Structure of GPR61-G protein complex stabilized by scFv16
Method: single particle / : Lees JA, Dias JM, Han S

EMDB-41145: 
Cryo-EM Structure of GPR61-
Method: single particle / : Lees JA, Dias JM, Han S

PDB-8tb0: 
Cryo-EM Structure of GPR61-G protein complex stabilized by scFv16
Method: single particle / : Lees JA, Dias JM, Han S

EMDB-28891: 
cryo-EM structure of a structurally designed Human metapneumovirus F protein in complex with antibody MPE8
Method: single particle / : Zhou T, Kwong PD, Morano NC, Ou L

PDB-8f6x: 
cryo-EM structure of a structurally designed Human metapneumovirus F protein in complex with antibody MPE8
Method: single particle / : Zhou T, Kwong PD, Morano NC, Ou L

EMDB-29725: 
Vaccine-elicited human antibody 2C06 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Morano NC, Shapiro L, Kwong PD

EMDB-29731: 
Vaccine-elicited human antibody 2C09 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD

PDB-8g4m: 
Vaccine-elicited human antibody 2C06 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Morano NC, Shapiro L, Kwong PD

PDB-8g4t: 
Vaccine-elicited human antibody 2C09 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD

EMDB-15786: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli (Apo form)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15787: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15788: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE (C387S mutant)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15789: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Lyso-PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15790: 
Cryo-EM structure apolipoprotein N-acyltransferase Lnt from E.coli in complex with FP3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

EMDB-15791: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Pam3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0k: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli (Apo form)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0l: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0m: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE (C387S mutant)
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0n: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Lyso-PE
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0o: 
Cryo-EM structure apolipoprotein N-acyltransferase Lnt from E.coli in complex with FP3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M

PDB-8b0p: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Pam3
Method: single particle / : Degtjarik O, Smithers L, Boland C, Caffrey M, Shalev Benami M
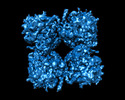
EMDB-29310: 
Deconvolved phiPA3 PhuN Tetramer, p2
Method: single particle / : Nieweglowska ES, Brilot AF, Mendez-Moran M, Kokontis C, Baek M, Li J, Cheng Y, Baker D, Bondy-Denomy J, Agard DA
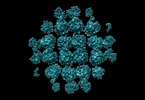
EMDB-29451: 
Tracing p2 phiPA3 PhuN Tetramer Interfaces
Method: single particle / : Nieweglowska ES, Brilot AF, Mendez-Moran M, Kokontis C, Baek M, Li J, Cheng Y, Baker D, Bondy-Denomy J, Agard DA

EMDB-29550: 
phiPA3 PhuN Tetramer, p2
Method: single particle / : Nieweglowska ES, Brilot AF, Mendez-Moran M, Kokontis C, Baek M, Li J, Cheng Y, Baker D, Bondy-Denomy J, Agard DA

PDB-8fne: 
phiPA3 PhuN Tetramer, p2
Method: single particle / : Nieweglowska ES, Brilot AF, Mendez-Moran M, Kokontis C, Baek M, Li J, Cheng Y, Baker D, Bondy-Denomy J, Agard DA

PDB-8fv5: 
Representation of 16-mer phiPA3 PhuN Lattice, p2
Method: single particle / : Nieweglowska ES, Brilot AF, Mendez-Moran M, Kokontis C, Baek M, Li J, Cheng Y, Baker D, Bondy-Denomy J, Agard DA

EMDB-13485: 
Cryo-EM structure of Bestrhodopsin (rhodopsin-rhodopsin-bestrophin) complex
Method: single particle / : Matzov D, Kaczmarczyk I, Shalev-Benami M

PDB-7pl9: 
Cryo-EM structure of Bestrhodopsin (rhodopsin-rhodopsin-bestrophin) complex
Method: single particle / : Matzov D, Kaczmarczyk I, Shalev-Benami M

EMDB-23835: 
Structure of p97 with substrate engaged
Method: single particle / : Xu Y, Han H, Cooney I, Hill CP, Shen PS

EMDB-26654: 
Wild Type p97-p47 complex (symmetric hexamer)
Method: single particle / : Xu Y, Han H, Cooney I, Shen PS, Hill CP

PDB-7mhs: 
Structure of p97 (subunits A to E) with substrate engaged
Method: single particle / : Xu Y, Han H, Cooney I, Hill CP, Shen PS

EMDB-25146: 
Broadly Neutralizing Antibody 10-40 in complex with prefusion SARS-CoV-2 B.1.351 Spike glycoprotein
Method: single particle / : Casner RG, Reddem ER, Shapiro L

EMDB-25166: 
Antibody 10-28 in complex with prefusion SARS-CoV-2 B.1.351 Spike glycoprotein
Method: single particle / : Casner RG, Shapiro L

EMDB-25167: 
Antibody 11-11 in complex with prefusion SARS-CoV-2 B.1.351 Spike glycoprotein
Method: single particle / : Casner RG, Shapiro L

EMDB-25076: 
LPHN3 (ADGRL3) 7TM domain bound to tethered agonist in complex with G protein heterotrimer
Method: single particle / : Barros-Alvarez X, Panova O, Skiniotis G

EMDB-25077: 
GPR56 (ADGRG1) 7TM domain bound to tethered agonist in complex with G protein heterotrimer
Method: single particle / : Barros-Alvarez X, Panova O, Skiniotis G

PDB-7sf7: 
LPHN3 (ADGRL3) 7TM domain bound to tethered agonist in complex with G protein heterotrimer
Method: single particle / : Barros-Alvarez X, Panova O, Skiniotis G

PDB-7sf8: 
GPR56 (ADGRG1) 7TM domain bound to tethered agonist in complex with G protein heterotrimer
Method: single particle / : Barros-Alvarez X, Panova O, Skiniotis G

EMDB-11927: 
Melanocortin receptor 4 (MC4R) Gs protein complex
Method: single particle / : Degtjarik O, Israeli H, Prabahar V, Shalev-Benami M

PDB-7aue: 
Melanocortin receptor 4 (MC4R) Gs protein complex
Method: single particle / : Degtjarik O, Israeli H, Prabahar V, Shalev-Benami M

EMDB-21389: 
Negative Staining Reconstruction of Tryptase Tetramer bound to E104 Fab
Method: single particle / : Koerber JT, Lazarus B, Maun H, Estevez A, Ciferri C

EMDB-11232: 
Cryo-EM structure of the highly atypical cytoplasmic ribosome of Euglena gracilis
Method: single particle / : Matzov D, Halfon H

PDB-6zj3: 
Cryo-EM structure of the highly atypical cytoplasmic ribosome of Euglena gracilis
Method: single particle / : Matzov D, Halfon H, Zimmerman E, Rozenberg H, Bashan A, Gray MW, Yonath AE, Shalev-Benami M

EMDB-10223: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A

EMDB-10224: 
Cryo-EM Structure of T. kodakarensis 70S ribosome in TkNat10 deleted strain
Method: single particle / : Matzov D, Sas-Chen A

EMDB-10503: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A

PDB-6skf: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A, Thomas JM, Santangelo T, Meier JL, Schwartz S, Shalev-Benami M

PDB-6skg: 
Cryo-EM Structure of T. kodakarensis 70S ribosome in TkNat10 deleted strain
Method: single particle / : Matzov D, Sas-Chen A, Thomas JM, Santangelo T, Meier JL, Schwartz S, Shalev-Benami M

PDB-6th6: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A, Thomas JM, Santangelo T, Meier JL, Schwartz S, Shalev-Benami M
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
