-Search query
-Search result
Showing 1 - 50 of 73 items for (author: maruyama & y)

EMDB-28092: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-093
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28090: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-040
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28091: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-045
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28093: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-156
Method: single particle / : Shek J, Callaway H, Li H, Yu X, Saphire EO

EMDB-28094: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-234
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28095: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-260
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28096: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-279
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28097: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-290
Method: single particle / : Yu X, Callaway H, Li H, Shek J, Saphire EO
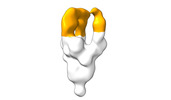
EMDB-28098: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-294
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28099: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-295
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28100: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-299
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28102: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-334
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
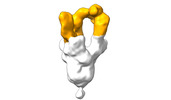
EMDB-28103: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-360
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28104: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-361
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28105: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-362
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28106: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-368
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28168: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-292
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28169: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-333
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
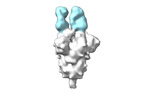
EMDB-28170: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-355
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28171: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-371
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-30862: 
Unliganded EGFR averaged cluster 5
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30864: 
Unliganded EGFR averaged cluster 21
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30714: 
Unliganded EGFR average cluster 1
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama I

EMDB-30715: 
Unliganded EGFR averaged cluster 2
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30716: 
Unliganded EGFR averaged cluster 3
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30717: 
Unliganded EGFR averaged cluster 4
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30719: 
Unliganded EGFR averaged cluster 6
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30721: 
Unliganded EGFR averaged cluster 7
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30722: 
Unliganded EGFR averaged cluster 9
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30723: 
Unliganded EGFR averaged cluster 8
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30724: 
Unliganded EGFR averaged cluster 9
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30725: 
Unliganded EGFR averaged cluster 10
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30726: 
Unliganded EGFR averaged cluster 11
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30727: 
Unliganded EGFR averaged cluster 12
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30728: 
Unliganded EGFR averaged cluster 13
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30729: 
Unliganded EGFR averaged cluster 14
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30730: 
Unliganded EGFR averaged cluster 15
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30731: 
Unliganded EGFR averaged cluster 16
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30732: 
Unliganded EGFR averaged cluster 17
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30733: 
Unliganded EGFR averaged cluster 18
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30734: 
Unliganded EGFR averaged cluster 19
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30735: 
Unliganded EGFR averaged cluster 20
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30737: 
Unliganded EGFR averaged cluster 22
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30738: 
Unliganded EGFR averaged cluster 23
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30739: 
Unliganded EGFR averaged cluster 24
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30740: 
Unliganded EGFR averaged cluster 25
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30741: 
Liganded EGFR averaged cluster 1
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30742: 
Liganded EGFR averaged cluster 2
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30743: 
Liganded EGFR averaged cluster 3
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN

EMDB-30744: 
Liganded EGFR averaged cluster 4
Method: electron tomography / : Purba ER, Saita EI, Akhouri RR, Ofverstedt LG, Wilken G, Skoglund U, Maruyama IN
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
