-Search query
-Search result
Showing all 26 items for (author: markl & j)
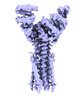
EMDB-50106: 
Artificial membrane protein TMHC4_R (ROCKET)
Method: single particle / : Abramsson ML, Anden O, Howard RJ, Lindahl E, Landreh M
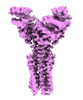
EMDB-50107: 
Artificial membrane protein TMHC4_R (ROCKET) mutant R9A/K10A/R13A
Method: single particle / : Abramsson M, Anden O, Howard RJ, Lindahl E, Landreh M

EMDB-19250: 
Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-19251: 
Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

PDB-8rjk: 
Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

PDB-8rjl: 
Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-16004: 
Structure of hexameric subcomplexes (Truncation Delta2-6) of the fractal citrate synthase from Synechococcus elongatus PCC7942
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-15529: 
Structure of a first level Sierpinski triangle formed by a citrate synthase
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-3717: 
Structure of differently sized plant protein
Method: single particle / : Saur M, Hennig R, Young P, Rusitzka K, Hellmann N, Heidrich J, Morgner N, Markl J, Schneider D

EMDB-3737: 
Structure of IM30 in C10-symmetrical assembly
Method: single particle / : Saur M, Hennig R, Young P, Rusitzka K, Hellmann N, Heidrich J, Morgner N, Markl J, Schneider D

EMDB-3738: 
Structure of IM30 in C12-symmetrical assembly
Method: single particle / : Saur M, Hennig R, Young P, Rusitzka K, Hellmann N, Heidrich J, Morgner N, Markl J, Schneider D

EMDB-3739: 
Structure of IM30 in C13-symmetrical assembly
Method: single particle / : Saur M, Hennig R, Young P, Rusitzka K, Hellmann N, Heidrich J, Morgner N, Markl J, Schneider D

EMDB-3740: 
Structure of IM30 in C14-symmetrical assembly
Method: single particle / : Saur M, Hennig R, Young P, Rusitzka K, Hellmann N, Heidrich J, Morgner N, Markl J, Schneider D
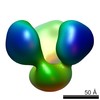
EMDB-2657: 
TEM of uncleaved soluble HIV envelope proteins
Method: single particle / : Arnold P, Himmels P, Weiss S, Decker TM, Markl J, Gatterdam V, Tampe R, Bartholomeaus P, Dietrich U, Duerr R

EMDB-6185: 
Electron cryo-microscopy of Melanoides tuberculata mega-hemocyanin
Method: single particle / : Gatsogiannis C, Hofnagel O, Markl J, Raunser S

EMDB-6186: 
Electron cryo-microscopy of Terebralia palustris mega-hemocyanin
Method: single particle / : Gatsogiannis C, Hofnagel O, Markl J, Raunser S

EMDB-2659: 
TEM of uncleaved soluble HIV envelope proteins
Method: single particle / : Arnold P, Himmels P, Weiss S, Decker TM, Markl J, Gatterdam V, Tampe R, Bartholomeaus P, Dietrich U, Duerr R

EMDB-2320: 
CryoEM Structure of keyhole limpet hemocyanin isoform 2 (KLH2)
Method: single particle / : Gatsogiannis C, Schnittger E, Markl SJ, Depoix F, Markl J

PDB-4bed: 
Keyhole limpet hemocyanin (KLH): 9A cryoEM structure and molecular model of the KLH1 didecamer reveal the interfaces and intricate topology of the 160 functional units
Method: single particle / : Gatsogiannis C, Markl J

EMDB-2055: 
Acetylcholine-binding protein in the hemolymph of the planorbid snail Biomphalaria glabrata is a pentagonal dodecahedron (60 subunits)
Method: single particle / : Saur M, Moeller V, Kapetanopoulos K, Braukmann S, Gebauer W, Tenzer S, Markl J

PDB-4aod: 
Biomphalaria glabrata Acetylcholine-binding protein type 1 (BgAChBP1)
Method: single particle / : Saur M, Moeller V, Kapetanopoulos K, Braukmann S, Gebauer W, Tenzer S, Markl J

PDB-4aoe: 
Biomphalaria glabrata Acetylcholine-binding protein type 2 (BgAChBP2)
Method: single particle / : Saur M, Moeller V, Kapetanopoulos K, Braukmann S, Gebauer W, Tenzer S, Markl J

EMDB-1434: 
Nautilus pompilius hemocyanin: 9 A cryo-EM structure and molecular model reveal the subunit pathway and the interfaces between the 70 functional units.
Method: single particle / : Gatsogiannis C, Moeller A, Depoix F, Meissner U, Markl J

EMDB-1628: 
8-A cryoEM structure and molecular model of the myriapod (Scutigera) 6x6mer hemocyanin: understanding a giant oxygen transport protein
Method: single particle / : Markl J, Moeller A, Martin AG, Rheinbay J, Gebauer W, Depoix F

EMDB-1304: 
Limulus polyphemus hemocyanin: 10 A cryo-EM structure, sequence analysis, molecular modelling and rigid-body fitting reveal the interfaces between the eight hexamers.
Method: single particle / : Martin AG, Depoix F, Stohr M, Meissner U, Hagner-Holler S, Hammouti K, Burmester T, Heyd J, Wriggers W, Markl J

EMDB-1569: 
Keyhole limpet hemocyanin (KLH): 9A cryoEM structure and molecular model of the didecamer reveal the interfaces and intricate topology of the 160 functional units
Method: single particle / : Gatsogiannis C, Markl J
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
