-Search query
-Search result
Showing all 46 items for (author: josé & m & carazo)

EMDB-23046: 
Cryo-EM structure of Mal de Rio Cuarto virus P9-1 viroplasm protein (decamer)
Method: single particle / : Llauger G, Melero R

EMDB-23047: 
Cryo-EM structure of Mal de Rio Cuarto virus P9-1 viroplasm protein (dodecamer)
Method: single particle / : Llauger G, Melero R

PDB-7kvc: 
Cryo-EM structure of Mal de Rio Cuarto virus P9-1 viroplasm protein (decamer)
Method: single particle / : Llauger G, Melero R, Monti D, Sycz G, Huck-Iriart C, Cerutti ML, Klinke S, Arranz R, Carazo JM, Goldbaum FA, del Vas M, Otero LH

PDB-7kvd: 
Cryo-EM structure of Mal de Rio Cuarto virus P9-1 viroplasm protein (dodecamer)
Method: single particle / : Llauger G, Melero R, Monti D, Sycz G, Huck-Iriart C, Cerutti ML, Klinke S, Arranz R, Carazo JM, Goldbaum FA, del Vas M, Otero LH
EMDB-13916: 
SARS-CoV-2 S protein S:A222V + S:D614G mutant 1-up
Method: single particle / : Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Lopez-Redondo ML, Frances-Gomez C, Ruiz-Rodriguez P, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, LLacer JL, Carazo JM
EMDB-13917: 
SARS-CoV-2 S protein S:A222V + S:D614G mutant 2-up
Method: single particle / : Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Lopez-Redondo ML, Ruiz-Rodriguez P, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM
EMDB-13918: 
SARS-CoV-2 S protein S:A222V + S:D614G mutant 3-down
Method: single particle / : Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Lopez-Redondo ML, Ruiz-Rodriguez P, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM
EMDB-13919: 
SARS-CoV-2 S protein S:D614G mutant 1-up
Method: single particle / : Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Lopez-Redondo ML, Frances-Gomez C, Ruiz-Rodriguez P, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, LLacer JL, Carazo JM
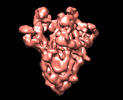
EMDB-13920: 
SARS-CoV-2 S protein S:D614G mutant 2-up
Method: single particle / : Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Lopez-Redondo ML, Ruiz-Rodriguez P, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM

PDB-7qdg: 
SARS-CoV-2 S protein S:A222V + S:D614G mutant 1-up
Method: single particle / : Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Lopez-Redondo ML, Frances-Gomez C, Ruiz-Rodriguez P, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, LLacer JL, Carazo JM

PDB-7qdh: 
SARS-CoV-2 S protein S:D614G mutant 1-up
Method: single particle / : Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Lopez-Redondo ML, Frances-Gomez C, Ruiz-Rodriguez P, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, LLacer JL, Carazo JM

EMDB-11831: 
BIP18SN. De novo coiled-coil-based bipyramidal protein
Method: single particle / : Lapenta F, Aupic J, Vezzoli M, Strmsek Z, Da Vela S, Svergun DI, Melero R, Carazo JM, Jerala R

EMDB-11328: 
SARS-CoV-2 spike in prefusion state
Method: single particle / : Martinez M, Marabini R, Carazo JM

EMDB-11336: 
SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up closed conformation)
Method: single particle / : Martinez M, Marabini R, Carazo JM

EMDB-11337: 
SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up open conformation)
Method: single particle / : Martinez M, Marabini R, Carazo JM

EMDB-11341: 
SARS-CoV-2 stabilized spike in prefusion state (1-up conformation)
Method: single particle / : Martinez M, Marabini R, Carazo JM

PDB-6zow: 
SARS-CoV-2 spike in prefusion state
Method: single particle / : Martinez M, Marabini R, Carazo JM

PDB-6zp5: 
SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up closed conformation)
Method: single particle / : Martinez M, Marabini R, Carazo JM

PDB-6zp7: 
SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up open conformation)
Method: single particle / : Martinez M, Marabini R, Carazo JM

EMDB-11008: 
Icosahedral reconstruction of human adenovirus D26 capsid
Method: single particle / : Huiskonen JT, Abrishami V

EMDB-11012: 
Localized reconstruction of human adenovirus D26 hexon type 3
Method: single particle / : Huiskonen JT, Abrishami V

EMDB-11009: 
Localized reconstruction of the trimeric fibre and its interactions with the penton base of human adenovirus HAdV-D26
Method: single particle / : Huiskonen JT, Abrishami V

EMDB-11010: 
Localized reconstruction of human adenovirus D26 hexon type 1
Method: single particle / : Huiskonen JT, Abrishami V

EMDB-11011: 
Localized reconstruction of human adenovirus D26 hexon type 2
Method: single particle / : Huiskonen JT, Abrishami V

EMDB-11013: 
Localized reconstruction of human adenovirus D26 hexon type 4
Method: single particle / : Huiskonen JT, Abrishami V

EMDB-4713: 
Negative staining of antibodies cooperative complex formed by human fab7B10 and fab2C1 bound to the N. meningitidis antigen factor H binding protein (fHbp)
Method: single particle / : Peschiera I, Ferlenghi I, Melero R, Liljeroos LJ, Paccagnini E, Giusti F, Carazo JM, Sorzano COS, Scarselli M

EMDB-4714: 
Negative staining of antibodies cooperative complex formed by human IgG1 mAb7B10 and mAb2C1 bound to the N. meningitidis antigen factor H binding protein (fHbp)
Method: single particle / : Peschiera I, Ferlenghi I, Melero R, Liljeroos LJ, Paccagnini E, Giusti F, Carazo JM, Sorzano COS, Scarselli M

EMDB-4715: 
Negative staining of antibodies cooperative complex formed by human IgG1 mAb1A3 and mAb1A12 bound to the N. meningitidis antigen factor H binding protein (fHbp)
Method: single particle / : Peschiera I, Ferlenghi I, Melero R, Liljeroos LJ, Paccagnini E, Giusti F, Carazo JM, Sorzano COS, Scarselli M

EMDB-3557: 
Centriolar Distal MT doublets in centrosome extracted from o.aries thymocytes
Method: subtomogram averaging / : Busselez J, Chichon FJ, Melero R, Carrascosa JL, Carazo JM

EMDB-3558: 
Centriolar Proximal MT triplet in centrosomes extracted from o.aries thymocytes
Method: subtomogram averaging / : Busselez J, Chichon FJ, Melero R, Carrascosa JL, Carazo JM

EMDB-3781: 
TET12SN density map (Design of in vivo self-assembling coiled-coil protein origami)
Method: single particle / : Melero R, Carazo JM

EMDB-3788: 
PYR16SN density map (Design of in vivo self-assembling coiled-coil protein origami)
Method: single particle / : Melero R, Carazo JM

EMDB-3789: 
TRIP18SN density map (Design of in vivo self-assembling coiled-coil protein origami)
Method: single particle / : Melero R, Carazo JM

EMDB-3540: 
Asymmetric reconstruction of MS2 virion
Method: single particle / : Koning RI, Gomez Blanco J, Akopjana I, Vargas J, Kazaks A, Tars K, Carazo JM, Koster AJ

EMDB-3402: 
Asymmetric cryo-EM reconstruction of phage MS2 reveals genome structure in situ
Method: single particle / : Koning RI, Gomez-Blanco J, Akopjana I, Vargas J, Kazaks A, Tars K, Carazo JM, Koster AJ

EMDB-3403: 
Asymmetric cryo-EM reconstruction of phage MS2 reveals genome structure in situ
Method: single particle / : Koning RI, Gomez-Blanco J, Akopjana I, Vargas J, Kazaks A, Tars K, Carazo JM, Koster AJ

EMDB-3404: 
Asymmetric cryo-EM reconstruction of phage MS2 reveals genome structure in situ
Method: single particle / : Koning RI, Gomez-Blanco J, Akopjana I, Vargas J, Kazaks A, Tars K, Carazo JM, Koster AJ

EMDB-3003: 
Structure of adenovirus light particle Ad5/FC31 L2
Method: single particle / : Condezo GN, Marabini R, Ayora S, Carazo JM, Chillon M, San Martin C

EMDB-3004: 
Structure of adenovirus light particle Ad5/FC31 L3
Method: single particle / : Condezo GN, Marabini R, Ayora S, Carazo JM, Chillon M, San Martin C

EMDB-1321: 
Quasi-atomic model of bacteriophage t7 procapsid shell: insights into the structure and evolution of a basic fold.
Method: single particle / : Agirrezabala X, Velazquez-Muriel J, Gomez-Puertas P, Scheres S, Carazo JM, Carrascosa JL

EMDB-1225: 
Loading a ring: structure of the Bacillus subtilis DnaB protein, a co-loader of the replicative helicase.
Method: single particle / : Nunez-Ramirez R, Velten M, Rivas G, Polard P, Carazo JM, Donate LE

EMDB-1134: 
Polymorphism and double hexamer structure in the archaeal minichromosome maintenance (MCM) helicase from Methanobacterium thermoautotrophicum.
Method: single particle / : Gomez-Llorente Y, Fletcher RJ, Chen XS, Carazo JM, San Martin C
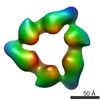
EMDB-1146: 
Quaternary polymorphism of replicative helicase G40P: structural mapping and domain rearrangement.
Method: single particle / : Nunez-Ramirez R, Robledo Y, mesa P, Alonso JC
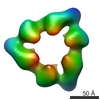
EMDB-1147: 
Quaternary polymorphism of replicative helicase G40P: structural mapping and domain rearrangement.
Method: single particle / : Nunez-Ramirez R, Robledo Y, mesa P, Alonso JC
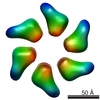
EMDB-1148: 
Quaternary polymorphism of replicative helicase G40P: structural mapping and domain rearrangement.
Method: single particle / : Nunez-Ramirez R, Robledo Y, mesa P, Alonso JC
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
