-Search query
-Search result
Showing 1 - 50 of 585 items for (author: hsu & a)

EMDB-39126: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly
Method: single particle / : Lin SC, Yang CY

EMDB-39127: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly
Method: single particle / : Lin SC, Yang CY

PDB-8ybx: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly
Method: single particle / : Lin SC, Yang CY

EMDB-37963: 
Cryo-EM structure of the hamster prion 23-144 fibril at pH 3.7
Method: helical / : Lee CH, Saw JE, Chen E, Wang CH, Chen R

PDB-8wzx: 
Cryo-EM structure of the hamster prion 23-144 fibril at pH 3.7
Method: helical / : Lee CH, Saw JE, Chen E, Wang CH, Chen R

EMDB-18334: 
Cryo-EM structure of the inward-facing FLVCR1
Method: single particle / : Weng TH, Wu D, Safarian S

EMDB-18335: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1
Method: single particle / : Weng TH, Wu D, Safarian S

EMDB-18336: 
Cryo-EM structure of the inward-facing FLVCR2
Method: single particle / : Weng TH, Wu D, Safarian S
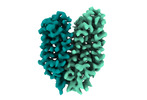
EMDB-18337: 
Cryo-EM structure of the outward-facing FLVCR2
Method: single particle / : Weng TH, Wu D, Safarian S

EMDB-18339: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2
Method: single particle / : Weng TH, Wu D, Safarian S

EMDB-19009: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1
Method: single particle / : Weng TH, Wu D, Safarian S

PDB-8qcs: 
Cryo-EM structure of the inward-facing FLVCR1
Method: single particle / : Weng TH, Wu D, Safarian S

PDB-8qct: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1
Method: single particle / : Weng TH, Wu D, Safarian S

PDB-8qcx: 
Cryo-EM structure of the inward-facing FLVCR2
Method: single particle / : Weng TH, Wu D, Safarian S

PDB-8qcy: 
Cryo-EM structure of the outward-facing FLVCR2
Method: single particle / : Weng TH, Wu D, Safarian S

PDB-8qd0: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2
Method: single particle / : Weng TH, Wu D, Safarian S

PDB-8r8t: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1
Method: single particle / : Weng TH, Wu D, Safarian S

EMDB-38691: 
Thiamine-bound human SLC19A3
Method: single particle / : Dang Y, Wang GP, Zhang Z

EMDB-38692: 
Pyridoxamine-bound human SLC19A3
Method: single particle / : Dang Y, Wang GP, Zhang Z

EMDB-38693: 
Fedratinib-bound human SLC19A3
Method: single particle / : Dang Y, Wang GP, Zhang Z
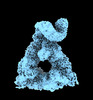
EMDB-40846: 
The C-terminal protease CtpA-LbcA complex of pseudomonas aeruginosa with the TPR at the low position
Method: single particle / : Hsu HC, Li H
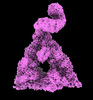
EMDB-40847: 
The C-terminal protease CtpA-LbcA complex of pseudomonas aeruginosa with the TPR at the high position
Method: single particle / : Hsu HC, Li H

EMDB-40848: 
The structure of C-terminal protease CtpA-LbcA complex of Pseudomonas aeruginosa
Method: single particle / : Hsu HC, Li H

EMDB-40849: 
Structure of the C-terminal protease CtpA-LbcA complex of Pseudomonas aeruginosa
Method: single particle / : Hsu HC, Li H

EMDB-40850: 
The C-terminal protease CtpA-LbcA complex of pseudomonas aeruginosa with the TPR at the high position
Method: single particle / : Hsu HC, Li H
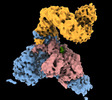
EMDB-40851: 
The C-terminal protease CtpA-LbcA complex of pseudomonas aeruginosa with the TPR at the low position
Method: single particle / : Hsu HC, Li H

EMDB-40852: 
Structure of the C-terminal protease CtpA-LbcA complex of Pseudomonas aeruginosa
Method: single particle / : Hsu HC, Li H

PDB-8sxe: 
Structure of the C-terminal protease CtpA-LbcA complex of Pseudomonas aeruginosa
Method: single particle / : Hsu HC, Li H

PDB-8sxf: 
The C-terminal protease CtpA-LbcA complex of pseudomonas aeruginosa with the TPR at the high position
Method: single particle / : Hsu HC, Li H

PDB-8sxg: 
The C-terminal protease CtpA-LbcA complex of pseudomonas aeruginosa with the TPR at the low position
Method: single particle / : Hsu HC, Li H

PDB-8sxh: 
Structure of the C-terminal protease CtpA-LbcA complex of Pseudomonas aeruginosa
Method: single particle / : Hsu HC, Li H

EMDB-38650: 
Additional map for SARS-CoV-2 Spike D614G variant, one RBD-up conformation 1 (PDB ID: 7EAZ; EMD-31047). Map was generated from heterogeneous refinement with downsampling in CryoSPARC
Method: single particle / : Yang TJ, Yu PY, Hsu STD

EMDB-29764: 
Structure of the Plasmodium falciparum 20S proteasome complexed with inhibitor TDI-8304
Method: single particle / : Hsu HC, Li H

EMDB-29765: 
Structure of the Plasmodium falciparum 20S proteasome beta-6 A117D mutant complexed with inhibitor WLW-vs
Method: single particle / : Hsu HC, Li H

EMDB-42148: 
Structure of human constitutive 20S proteasome complexed with the inhibitor TDI-8304
Method: single particle / : Hsu HC, Li H

PDB-8g6e: 
Structure of the Plasmodium falciparum 20S proteasome complexed with inhibitor TDI-8304
Method: single particle / : Hsu HC, Li H

PDB-8g6f: 
Structure of the Plasmodium falciparum 20S proteasome beta-6 A117D mutant complexed with inhibitor WLW-vs
Method: single particle / : Hsu HC, Li H

PDB-8ud9: 
Structure of human constitutive 20S proteasome complexed with the inhibitor TDI-8304
Method: single particle / : Hsu HC, Li H

EMDB-29172: 
Cryo-EM structure of Cryptococcus neoformans trehalose-6-phosphate synthase homotetramer in complex with uridine diphosphate and glucose-6-phosphate
Method: single particle / : Washington EJ, Brennan RG

PDB-8fhw: 
Cryo-EM structure of Cryptococcus neoformans trehalose-6-phosphate synthase homotetramer in complex with uridine diphosphate and glucose-6-phosphate
Method: single particle / : Washington EJ, Brennan RG

EMDB-42589: 
Prototypic SARS-CoV-2 spike (containing K417) in the closed conformation
Method: single particle / : Geng Q, Liu B, Li F

EMDB-42590: 
Prototypic SARS-CoV-2 spike (containing K417) in the open conformation
Method: single particle / : Geng Q, Liu B, Li F

EMDB-42591: 
Prototypic SARS-CoV-2 spike (containing V417) in the closed conformation
Method: single particle / : Geng Q, Liu B, Li F

EMDB-42592: 
Prototypic SARS-CoV-2 spike (containing V417) in the open conformation
Method: single particle / : Geng Q, Liu B, Li F

PDB-8uul: 
Prototypic SARS-CoV-2 spike (containing K417) in the closed conformation
Method: single particle / : Geng Q, Liu B, Li F

PDB-8uum: 
Prototypic SARS-CoV-2 spike (containing K417) in the open conformation
Method: single particle / : Geng Q, Liu B, Li F

PDB-8uun: 
Prototypic SARS-CoV-2 spike (containing V417) in the closed conformation
Method: single particle / : Geng Q, Liu B, Li F
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search






 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
