-Search query
-Search result
Showing all 47 items for (author: hodes & a)

EMDB-31806: 
Telomeric mononucleosome
Method: single particle / : Soman A

EMDB-31810: 
Telomeric Dinucleosome
Method: single particle / : Soman A

EMDB-31815: 
Telomeric Dinucleosome in open state
Method: single particle / : Soman A

EMDB-31816: 
Telomeric trinucleosome
Method: single particle / : Soman A

EMDB-31823: 
Telomeric tetranucleosome
Method: single particle / : Soman A

EMDB-31826: 
Telomeric trinucleosome in open state
Method: single particle / : Soman A

EMDB-31832: 
Telomeric tetranucleosome in open state
Method: single particle / : Soman A

EMDB-31907: 
Telomeric Dinucleosome at 4.6 angstroms
Method: single particle / : Soman A

EMDB-31908: 
Telomeric Dinucleosome at 5angstroms
Method: single particle / : Soman A

EMDB-31909: 
Open dinucleosome at 6.6 angstroms
Method: single particle / : Soman A
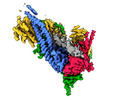
EMDB-13785: 
Cryo-EM map of clamped S.cerevisiae condensin-DNA complex (pentamer dataset)
Method: single particle / : Lee BG, Rhodes J, Lowe J
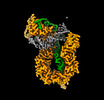
EMDB-13786: 
Cryo-EM structure of S.cerevisiae condensin Ycg1-Brn1-DNA complex
Method: single particle / : Lee BG, Rhodes J

EMDB-13787: 
Cryo-EM map of clamped S.cerevisiae condensin complex on circular DNA
Method: single particle / : Lee BG, Rhodes J, Lowe J
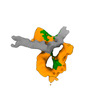
EMDB-13788: 
Cryo-EM map of S.cerevisiae condensin Ycg1-Brn1 complex on circular DNA
Method: single particle / : Lee BG, Rhodes J, Lowe J

PDB-7q2z: 
Cryo-EM structure of S.cerevisiae condensin Ycg1-Brn1-DNA complex
Method: single particle / : Lee BG, Rhodes J, Lowe J

EMDB-13783: 
Cryo-EM structure of clamped S.cerevisiae condensin-DNA complex (Form I)
Method: single particle / : Lee BG, Rhodes J, Lowe J

EMDB-13784: 
Cryo-EM structure of clamped S.cerevisiae condensin-DNA complex (form II)
Method: single particle / : Lee BG, Rhodes J, Lowe J

PDB-7q2x: 
Cryo-EM structure of clamped S.cerevisiae condensin-DNA complex (Form I)
Method: single particle / : Lee BG, Rhodes J, Lowe J

PDB-7q2y: 
Cryo-EM structure of clamped S.cerevisiae condensin-DNA complex (form II)
Method: single particle / : Lee BG, Rhodes J, Lowe J

EMDB-30596: 
Cryo-EM map of POT1 bound by TPP1 in closed conformation
Method: single particle / : Smith EW, Liu ZB, Lattmann S, Ahsan B, Rhodes D

EMDB-30597: 
Cryo-EM map of POT1 bound by TPP1 in extended conformation
Method: single particle / : Smith EW, Liu ZB, Lattmann S, Ahsan B, Rhodes D

EMDB-4704: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 1, free nuclesome
Method: single particle / : Marabelli C, Pilotto S

EMDB-4705: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 2
Method: single particle / : Marabelli C, Pilotto S

EMDB-4710: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 3
Method: single particle / : Marabelli C, Pilotto S, Chittori S, Subramaniam S, Mattevi A

EMDB-4711: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 4
Method: single particle / : Marabelli C, Pilotto S, Chittori S, Subramaniam S, Mattevi A

EMDB-4712: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 5
Method: single particle / : Marabelli C, Pilotto S, Chittori S, Subramaniam S, Mattevi A

PDB-6r1t: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 1, free nuclesome
Method: single particle / : Marabelli C, Pilotto S, Chittori S, Subramaniam S, Mattevi A

PDB-6r1u: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 2
Method: single particle / : Marabelli C, Pilotto S, Chittori S, Subramaniam S, Mattevi A

PDB-6r25: 
Structure of LSD2/NPAC-linker/nucleosome core particle complex: Class 3
Method: single particle / : Marabelli C, Pilotto S, Chittori S, Subramaniam S, Mattevi A

EMDB-6291: 
16 Angstrom cryo-EM reconstruction of the alpha, beta, gamma TF55 chaperonin
Method: single particle / : Chaston JJ, Stewart AG, Smits C, Aragao D, Struwe W, Benesch J, Xwong A, Ling M, Ashsan B, Sandin S, Rhodes D, Molugu SK, Bernal RA, Stock D

EMDB-5764: 
A new topology of the HK97-like fold revealed in Bordetella bacteriophage by cryoEM at 3.5A resolution
Method: single particle / : Zhang X, Guo H, Jin L, Czornyj E, Hodes A, Hui WH, Nieh AW, Miller JF, Zhou ZH

EMDB-5765: 
A new topology of the HK97-like fold revealed in Bordetella bacteriophage by cryoEM at 3.5A resolution
Method: single particle / : Zhang X, Guo H, Jin L, Czornyj E, Hodes A, Hui WH, Nieh AW, Miller JF, Zhou ZH

EMDB-5766: 
A new topology of the HK97-like fold revealed in Bordetella bacteriophage by cryoEM at 3.5A resolution
Method: single particle / : Zhang X, Guo H, Jin L, Czornyj E, Hodes A, Hui WH, Nieh AW, Miller JF, Zhou ZH

PDB-3j4u: 
A new topology of the HK97-like fold revealed in Bordetella bacteriophage: non-covalent chainmail secured by jellyrolls
Method: single particle / : Zhang X, Guo H, Jin L, Czornyj E, Hodes A, Hui WH, Nieh AW, Miller JF, Zhou ZH

EMDB-2310: 
Three-dimensional structure of active, full-length human telomerase dimer, determined by single-particle electron microscopy in negative stain
Method: single particle / : Sauerwald A, Sandin S, Cristofari G, Scheres SHW, Lingner J, Rhodes D

EMDB-2311: 
Three-dimensional structure of active, full-length human telomerase. Independently refined open monomer structure, determined by single-particle electron microscopy in negative stain
Method: single particle / : Sauerwald A, Sandin S, Cristofari G, Scheres SHW, Lingner J, Rhodes D

EMDB-2312: 
Three-dimensional structure of active, full-length human telomerase. Independently refined closed monomer structure, determined by single-particle electron microscopy in negative stain
Method: single particle / : Sauerwald A, Sandin S, Cristofari G, Scheres SHW, Lingner J, Rhodes D

EMDB-5200: 
5.4-Angstrom cryoEM structure of the Bordetella Bacteriophage capsid
Method: single particle / : Jin L, Hodes A, Hui WH, Zhang X, Yu X, Miller JF, Zhou ZH
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search












 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
