[English] 日本語
 Yorodumi
Yorodumi- EMDB-1619: Three-dimensional Structure of Tropism-Switching Bordetella Bacte... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1619 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Three-dimensional Structure of Tropism-Switching Bordetella Bacteriophage | |||||||||
 Map data Map data | Bottom view of the BPP-1 phage particle (after average) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  cryo-electron tomography / Bordetella plus phage / major tropism determinant cryo-electron tomography / Bordetella plus phage / major tropism determinant | |||||||||
| Biological species |  Bordetella phage BPP-1 (virus) Bordetella phage BPP-1 (virus) | |||||||||
| Method | subtomogram averaging /  cryo EM / Resolution: 40.0 Å cryo EM / Resolution: 40.0 Å | |||||||||
 Authors Authors | Dai W / Hodes A / Hui WH / Gingery M / Miller JF / Zhou ZH | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2010 Journal: Proc Natl Acad Sci U S A / Year: 2010Title: Three-dimensional structure of tropism-switching Bordetella bacteriophage. Authors: Wei Dai / Asher Hodes / Wong H Hui / Mari Gingery / Jeff F Miller / Z Hong Zhou /  Abstract: Bacteriophage BPP-1, which infects Bordetella species, can switch its specificity by mutations to the ligand-binding surface of its major tropism-determinant protein, Mtd. This targeted mutagenesis ...Bacteriophage BPP-1, which infects Bordetella species, can switch its specificity by mutations to the ligand-binding surface of its major tropism-determinant protein, Mtd. This targeted mutagenesis results from the activity of a phage-encoded diversity-generating retroelement. Purified Mtd binds its receptor with low affinity, yet BPP-1 binding and infection of Bordettella cells are efficient because of high-avidity binding between phage-associated Mtd and its receptor. Here, using an integrative approach of three-dimensional (3D) structural analyses of the entire phage by cryo-electron tomography and single-prticle cryo-electron microscopy, we provide direct localization of Mtd in the phage and the structural basis of the high-avidity binding of the BPP-1 phage. Our structure shows that each BPP-1 particle has a T = 7 icosahedral head and an unusual tail apparatus consisting of a short central tail "hub," six short tail spikes, and six extended tail fibers. Subtomographic averaging of the tail fiber maps revealed a two-lobed globular structure at the distal end of each long tail fiber. Tomographic reconstructions of immuno-gold-labeled BPP-1 directly localized Mtd to these globular structures. Finally, our icosahedral reconstruction of the BPP-1 head at 7A resolution reveals an HK97-like major capsid protein stabilized by a smaller cementing protein. Our structure represents a unique bacteriophage reconstruction with its tail fibers and ligand-binding domains shown in relation to its tail apparatus. The localization of Mtd at the distal ends of the six tail fibers explains the high avidity binding of Mtd molecules to cell surfaces for initiation of infection. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1619.map.gz emd_1619.map.gz | 7.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1619-v30.xml emd-1619-v30.xml emd-1619.xml emd-1619.xml | 8.4 KB 8.4 KB | Display Display |  EMDB header EMDB header |
| Images |  1619.png 1619.png | 107.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1619 http://ftp.pdbj.org/pub/emdb/structures/EMD-1619 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1619 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1619 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1619.map.gz / Format: CCP4 / Size: 8.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1619.map.gz / Format: CCP4 / Size: 8.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Bottom view of the BPP-1 phage particle (after average) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.86 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
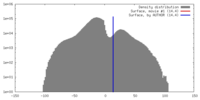
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bvg plus tropic phage-1 (BPP-1) bacteriophage
| Entire | Name: Bvg plus tropic phage-1 (BPP-1) bacteriophage |
|---|---|
| Components |
|
-Supramolecule #1000: Bvg plus tropic phage-1 (BPP-1) bacteriophage
| Supramolecule | Name: Bvg plus tropic phage-1 (BPP-1) bacteriophage / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Bordetella phage BPP-1
| Supramolecule | Name: Bordetella phage BPP-1 / type: virus / ID: 1 / Name.synonym: BPP-1 bacteriophage / NCBI-ID: 194699 / Sci species name: Bordetella phage BPP-1 / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: BPP-1 bacteriophage |
|---|---|
| Host (natural) | Organism:   Bordetella (bacteria) / synonym: BACTERIA(EUBACTERIA) Bordetella (bacteria) / synonym: BACTERIA(EUBACTERIA) |
| Virus shell | Shell ID: 1 / Diameter: 680 Å / T number (triangulation number): 7 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Grid | Details: 200 mesh grid |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER / Method: Blot for 1 second before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 38200 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2 mm / Nominal defocus max: 6.0 µm / Nominal defocus min: 4.0 µm / Nominal magnification: 23000 Bright-field microscopy / Cs: 2 mm / Nominal defocus max: 6.0 µm / Nominal defocus min: 4.0 µm / Nominal magnification: 23000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen-cooled cryo specimen holder Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: -70 ° / Tilt series - Axis1 - Max angle: 70 ° |
| Image recording | Category: CCD / Film or detector model: GENERIC CCD / Average electron dose: 1 e/Å2 |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 40.0 Å / Software - Name: ProTomo / Details: Final map was averaged from 60 particles |
|---|
 Movie
Movie Controller
Controller


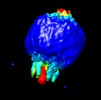







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)