-Search query
-Search result
Showing 1 - 50 of 68 items for (author: esser & l)

EMDB-18245: 
Plunge-frozen (control) map of beta-galactosidase
Method: single particle / : Esser TK, Boehning J, Bharat TAM, Rauschenbach S

EMDB-18586: 
Cryo-EM reconstruction of crosslinked native Bacillus subtilis collided disome binding MutS2 SMR and KOW domains
Method: single particle / : Park E, Mackens-Kiani T, Berhane R, Esser H, Erdenebat C, Burroughs AM, Berninghausen O, Aravind L, Beckmann R, Green R, Buskirk AR

EMDB-18585: 
Cryo-EM reconstruction of Bacillus subtilis collided disome binding MutS2 SMR and KOW domains
Method: single particle / : Park E, Mackens-Kiani T, Berhane R, Esser H, Erdenebat C, Burroughs AM, Berninghausen O, Aravind L, Beckmann R, Green R, Buskirk AR

EMDB-18901: 
Bacillus subtilis MutS2-collided disome complex (stalled 70S)
Method: single particle / : Park E, Mackens-Kiani T, Berhane R, Esser H, Erdenebat C, Burroughs AM, Berninghausen O, Aravind L, Beckmann R, Green R, Buskirk AR

PDB-8r55: 
Bacillus subtilis MutS2-collided disome complex (collided 70S)
Method: single particle / : Park E, Mackens-Kiani T, Berhane R, Esser H, Erdenebat C, Burroughs AM, Berninghausen O, Aravind L, Beckmann R, Green R, Buskirk AR

EMDB-18244: 
ESIBD structure of beta-galactosidase
Method: single particle / : Esser T, Boehning J, Bharat TAM, Rauschenbach S

PDB-8q7y: 
ESIBD structure of beta-galactosidase
Method: single particle / : Esser T, Boehning J, Bharat TAM, Rauschenbach S

EMDB-18558: 
Bacillus subtilis MutS2-collided disome complex (stalled 70S)
Method: single particle / : Park E, Mackens-Kiani T, Berhane R, Esser H, Erdenebat C, Burroughs AM, Berninghausen O, Aravind L, Beckmann R, Green R, Buskirk AR

PDB-8qpp: 
Bacillus subtilis MutS2-collided disome complex (stalled 70S)
Method: single particle / : Park E, Mackens-Kiani T, Berhane R, Esser H, Erdenebat C, Burroughs AM, Berninghausen O, Aravind L, Beckmann R, Green R, Buskirk AR

EMDB-17464: 
EM structure of endogenous TREX complex from S. cerevisiae
Method: single particle / : Kern C, Straesser K

EMDB-26203: 
Structure of mitochondrial bc1 in complex with ck-2-68
Method: single particle / : Xia D, Esser L

PDB-7tz6: 
Structure of mitochondrial bc1 in complex with ck-2-68
Method: single particle / : Xia D, Esser L, Zhou F, Huang R

EMDB-25989: 
Rhodobacter sphaeroides Mitochondrial respiratory chain complex
Method: single particle / : Xia D, Zhou F, Esser L, Huang R

PDB-7tlj: 
Rhodobacter sphaeroides Mitochondrial respiratory chain complex
Method: single particle / : Xia D, Zhou F, Esser L, Huang R

EMDB-26812: 
G. haemolysans IgA1 protease
Method: single particle / : Eisenmesser EZ, Zheng H

EMDB-26813: 
IgA1 Protease with IgA1 substrate
Method: single particle / : Eisenmesser ZE, Zheng H

PDB-7uvk: 
G. haemolysans IgA1 protease
Method: single particle / : Eisenmesser EZ, Zheng H

PDB-7uvl: 
IgA1 Protease with IgA1 substrate
Method: single particle / : Eisenmesser ZE, Zheng H

EMDB-27254: 
SARS-CoV-2 Spike RBD in complex with DMAbs 2130 and 2196
Method: single particle / : Du J, Cui J, Pallesen J

EMDB-27255: 
SARS-CoV-2 Spike RBD in complex with DMAb 2196
Method: single particle / : Du J, Cui J, Pallesen J

PDB-8d8q: 
SARS-CoV-2 Spike RBD in complex with DMAbs 2130 and 2196
Method: single particle / : Du J, Cui J, Pallesen J

PDB-8d8r: 
SARS-CoV-2 Spike RBD in complex with DMAb 2196
Method: single particle / : Du J, Cui J, Pallesen J

EMDB-25200: 
CryoEM structure of SARS-CoV-2 HexaPro spike in complex with S2-binding CV3-25
Method: single particle / : Chen Y, Huang R, Pazgier M

EMDB-12734: 
Cryo-EM structure of a Bacillus subtilis MifM-stalled ribosome-nascent chain complex with (p)ppGpp-SRP bound
Method: single particle / : Kratzat H, Czech L, Berninghausen O, Bange G, Beckmann R

EMDB-12735: 
Cryo-EM map of a Bacillus subtilis MifM-stalled ribosome-nascent chain complex with GMPPNP-SRP bound
Method: single particle / : Kratzat H, Berninghausen O, Beckmann R

EMDB-13839: 
Cryo-EM structure of an Escherichia coli TnaC-stalled FtsQ ribosome-nascent chain complex with GMPPNP-SRP bound
Method: single particle / : Esser HF, Kratzat H, Musial J, Berninghausen O, Beckmann R

EMDB-13840: 
Cryo-EM structure of an Escherichia coli TnaC-stalled FtsQ ribosome-nascent chain complex with (p)ppGpp-SRP bound
Method: single particle / : Esser HF, Kratzat H, Musial J, Berninghausen O, Beckmann R

PDB-7o5b: 
Cryo-EM structure of a Bacillus subtilis MifM-stalled ribosome-nascent chain complex with (p)ppGpp-SRP bound
Method: single particle / : Kratzat H, Czech L, Berninghausen O, Bange G, Beckmann R

EMDB-25564: 
Subtomogram averaging of SARS-CoV-2 Spike Protein bound to CV3-1
Method: subtomogram averaging / : Li W, Mothes W

EMDB-25565: 
Subtomogram averaging of SARS-CoV-2 Spike Protein bound to Fab CV3-25
Method: subtomogram averaging / : Li W, Mothes W

EMDB-25566: 
Subtomogram averaging of SARS-CoV-2 Spike Protein
Method: subtomogram averaging / : Li W, Mothes W

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
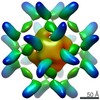
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-23211: 
Cryo-EM structure of human ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1
Method: single particle / : Lees JA, Han S

EMDB-23215: 
Cryo-EM structure of protein encoded by vaccine candidate BNT162b2
Method: single particle / : Lees JA, Han S

PDB-7l7f: 
Cryo-EM structure of human ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1
Method: single particle / : Lees JA, Han S

PDB-7l7k: 
Cryo-EM structure of protein encoded by vaccine candidate BNT162b2
Method: single particle / : Lees JA, Han S

EMDB-22204: 
Streptococcus Pneumoniae IgA1 Protease with IgA1 substrate
Method: single particle / : Eisenmesser EZ, Zheng H

PDB-6xja: 
Streptococcus Pneumoniae IgA1 Protease with IgA1 substrate
Method: single particle / : Eisenmesser EZ, Zheng H

EMDB-22205: 
IgA1 Protease
Method: single particle / : Eisenmesser EZ, Zheng H

EMDB-22328: 
IgA1 Protease in complex with neutralizing mAb
Method: single particle / : Eisenmesser EZ, Zheng H

PDB-6xjb: 
IgA1 Protease
Method: single particle / : Eisenmesser EZ, Zheng H

PDB-7jgj: 
IgA1 Protease in complex with neutralizing mAb
Method: single particle / : Eisenmesser EZ, Zheng H

EMDB-22913: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the ACE2 protein decoy, CTC-445.2 (State 1)
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22914: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the ACE2 protein decoy, CTC-445.2 (State 2)
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22915: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the ACE2 protein decoy, CTC-445.2 (State 4)
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-22916: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the ACE2 protein decoy, CTC-445.2 (State 4)
Method: single particle / : Barnes CO, Bjorkman PJ

PDB-7kl9: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the ACE2 protein decoy, CTC-445.2 (State 4)
Method: single particle / : Barnes CO, Bjorkman PJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
