-Search query
-Search result
Showing 1 - 50 of 173 items for (author: chait & b)

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-40699: 
SARS-CoV-2 replication-transcription complex bound to nsp9 and UMPCPP, as a pre-catalytic NMPylation intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA

EMDB-40707: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9, as a noncatalytic RNA-nsp9 binding mode
Method: single particle / : Small GI, Darst SA, Campbell EA

EMDB-40708: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9 and GDP-betaS, as a pre-catalytic deRNAylation/mRNA capping intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA

PDB-8sq9: 
SARS-CoV-2 replication-transcription complex bound to nsp9 and UMPCPP, as a pre-catalytic NMPylation intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA

PDB-8sqj: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9, as a noncatalytic RNA-nsp9 binding mode
Method: single particle / : Small GI, Darst SA, Campbell EA

PDB-8sqk: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9 and GDP-betaS, as a pre-catalytic deRNAylation/mRNA capping intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA
EMDB-41123: 
Composite multiscale structure of the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41114: 
Nuclear double outer ring of the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41116: 
Lumenal ring of the isolated yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41117: 
Pom34-Pom152 membrane attachment site yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41119: 
Cytoplasmic outer ring of the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41120: 
Central transporter/plug yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41121: 
Double outer ring connectors of the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP
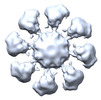

EMDB-41122: 
Connectors in cytoplasmic outer ring of yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41285: 
Double nuclear outer ring of Nup84-complexes from the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-41300: 
Inner spoke ring of the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

PDB-8t9l: 
Pom34-Pom152 membrane attachment site yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

PDB-8tie: 
Double nuclear outer ring of Nup84-complexes from the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

PDB-8tj5: 
Inner spoke ring of the yeast NPC
Method: single particle / : Akey CW, Echeverria I, Ouch C, Fernandez-Martinez J, Rout MP

EMDB-29910: 
SARS-CoV-2 Spike H655Y variant, One RBD Open
Method: single particle / : Egri SB, Shen K, Luban J

PDB-8gb0: 
SARS-CoV-2 Spike H655Y variant, One RBD Open
Method: single particle / : Egri SB, Shen K, Luban J

EMDB-26639: 
SARS-CoV-2 replication-transcription complex bound to Remdesivir triphosphate, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26641: 
SARS-CoV-2 replication-transcription complex bound to ATP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26642: 
SARS-CoV-2 replication-transcription complex bound to UTP, in a pre-catalytic state.
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26645: 
SARS-CoV-2 replication-transcription complex bound to GTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
